The London Calling 2024 Oxford Nanopore conference took place this week, and one of the last presentations was the tech update by Clive Brown.
Since the company is now in the stock market, the presentations always start with a Caution concerning forward-looking statements.
The first few comments were about the history of Oxford Nanopore, the core principles established very early on for the platform and what they meant for the products. One of the big parts was being able to put biological membranes on top of an ASIC, then make them stable enough that you could ship them around the world. Many people didn’t believe this to be possible, and over a series of iterations, which took more than 2 years, it became robust enough to start shipping the first batch of flowcells. There was lots of criticism back then (Clive used a different s*** word for criticism), but the company eventually succeeded in product number 1.
The MinION is how everything started, a product that, unlike all the other NGS sequencers out there, had no pumps, no fluidics, the only pumping was done by the customer’s thumb when pipetting the sample into the flowcell. Clive described the vision of this disruptive product, which was unlike any other NGS sequencer, and how, for the platform to succeed, everything else needed to be disruptive. Quite early in the process, when the company had only 100 employees, they came up with the first Early Access program, named MAP and shipped to about 400 customers. It was a difficult process, some customers managed to get good results, others couldn’t work it out, but from then on, ONT was on the map and got lots of feedback for their platform and their future plans.
London Calling is a type of conference mimicking what Apple used to do with their Apple Dev conference, and other equivalents in Silicon Valley: a way to talk to customers, a way to disseminate experiences using the technology, a way for the customers to influence the direction of the future products, etc. and that is what London Calling is.
The basics of the technology, where things are looking at the big picture, it took a lot of effort to get there. When the company presented early on, there were people taking notes in the audience, then they went and wrote patent applications to then be able to throw them back against ONT in a patent war that lasted years. At one point, there were 3 different large companies trying to sue ONT, but they’ve overcome those issues.
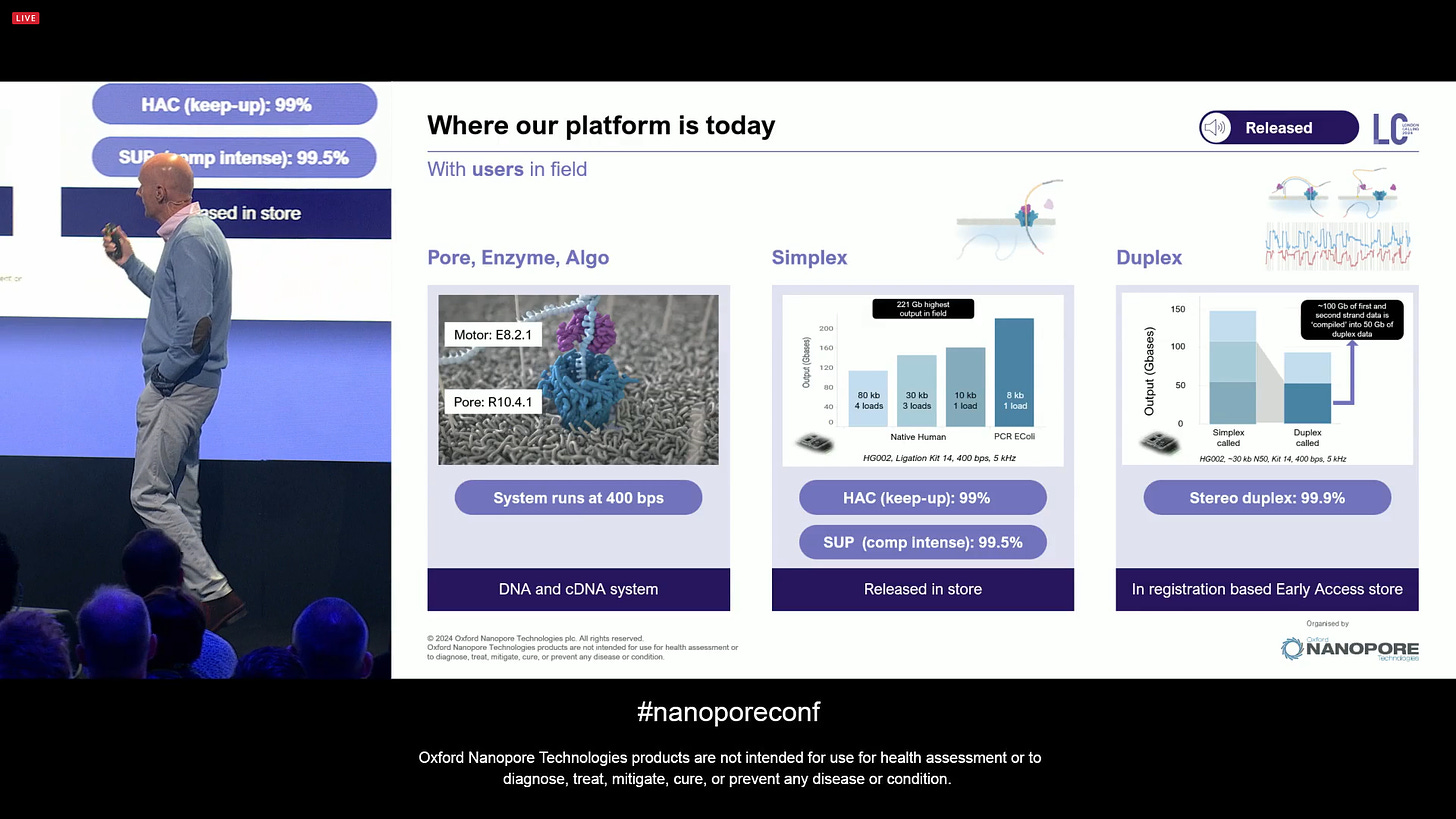
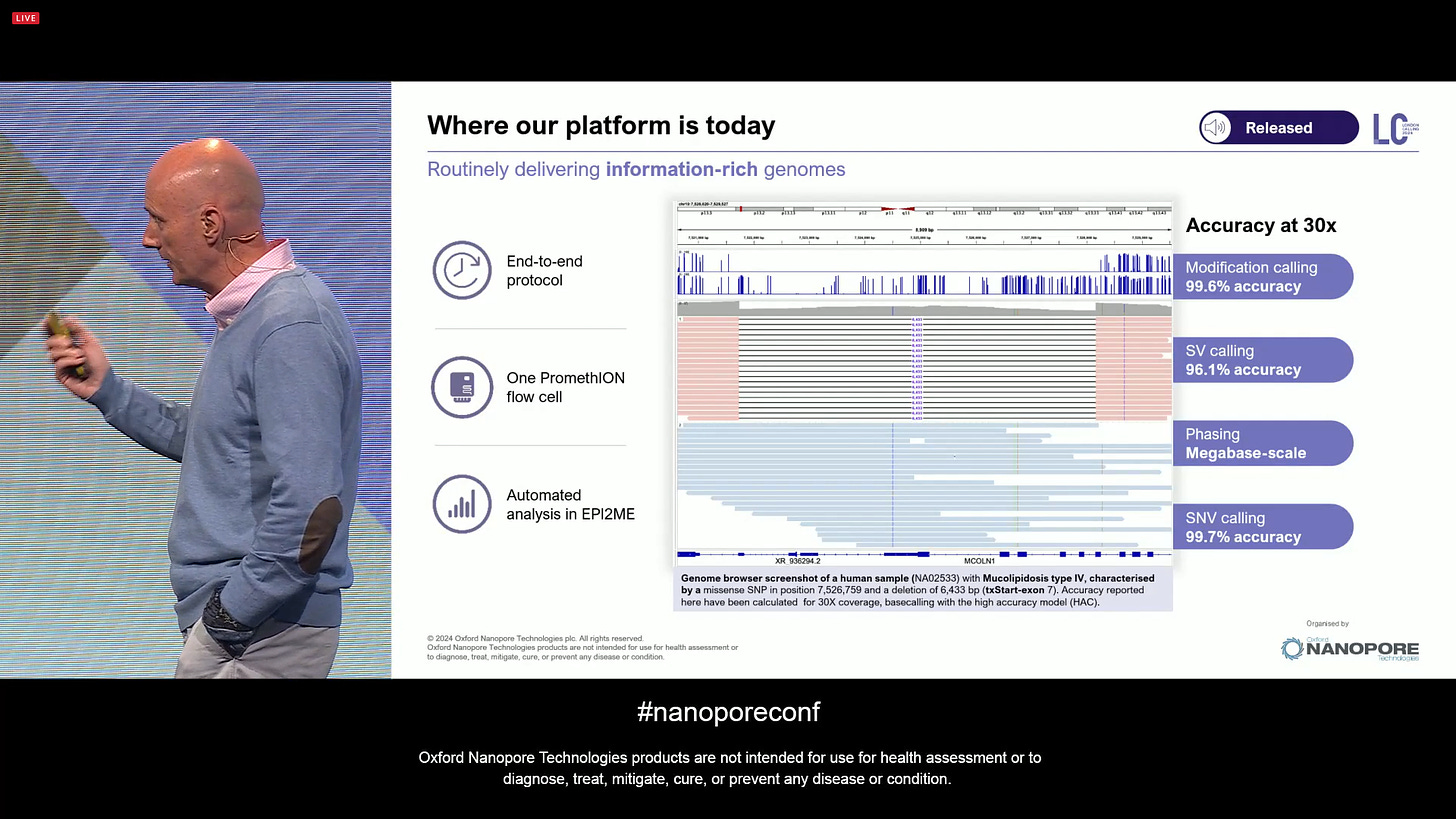
The technology platform as it is today, with some key elements: an End-to-end protocol, with the PromethION flowcells delivering the type of throughput that can be used for many types of applications, not only SNV and SV calling, but also Phasing, and Modification calling (epigenetic marks), and all with software tools including the Automated analysis in EPI2ME.
The device is still not lab free, but close to it. A lot of progress, enables a lot of experimental flexibility, which has seen the device been used by NASA in space, also in submarines, at the top of mountains, remote areas, all with minimal portable lab ancillary equipment.
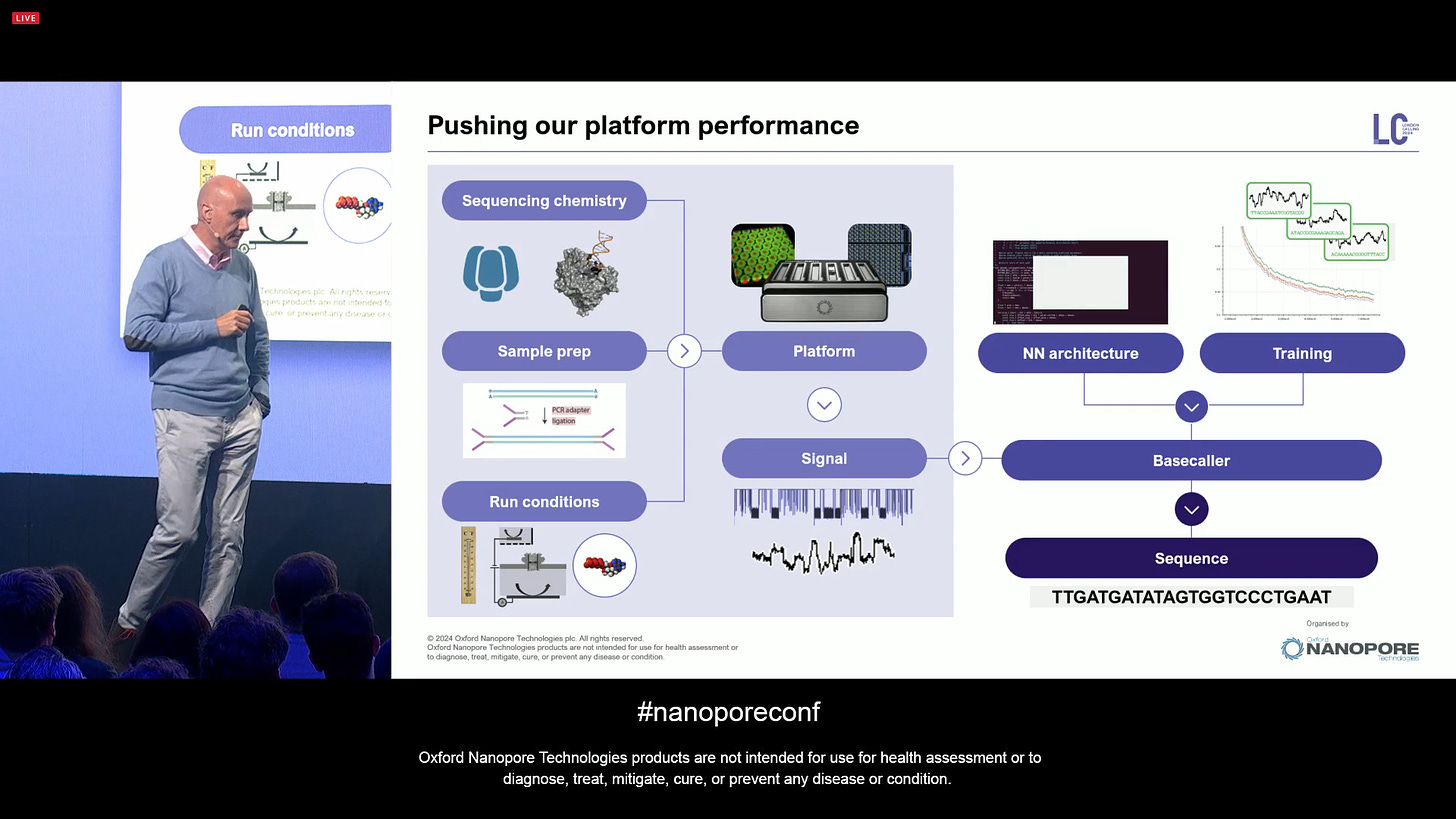
An early design choice of the platform is to build a lot of headroom, for example in the electronics. Making sure there would be flexibility in different orthogonal aspects of the technology, allowing for an iterative tuning in aspects like speed, the salts in the buffers, etc. There are many ways to improve the technology other than the scale, which is a reference to what Illumina ILMN 0.00%↑ has done in the last few years: make cheaper sequencing by making larger-scale sequencers, with the most expensive nowadays being the Illumina NovaSeqX which is $1.25M and the size of a fridge/freezer.
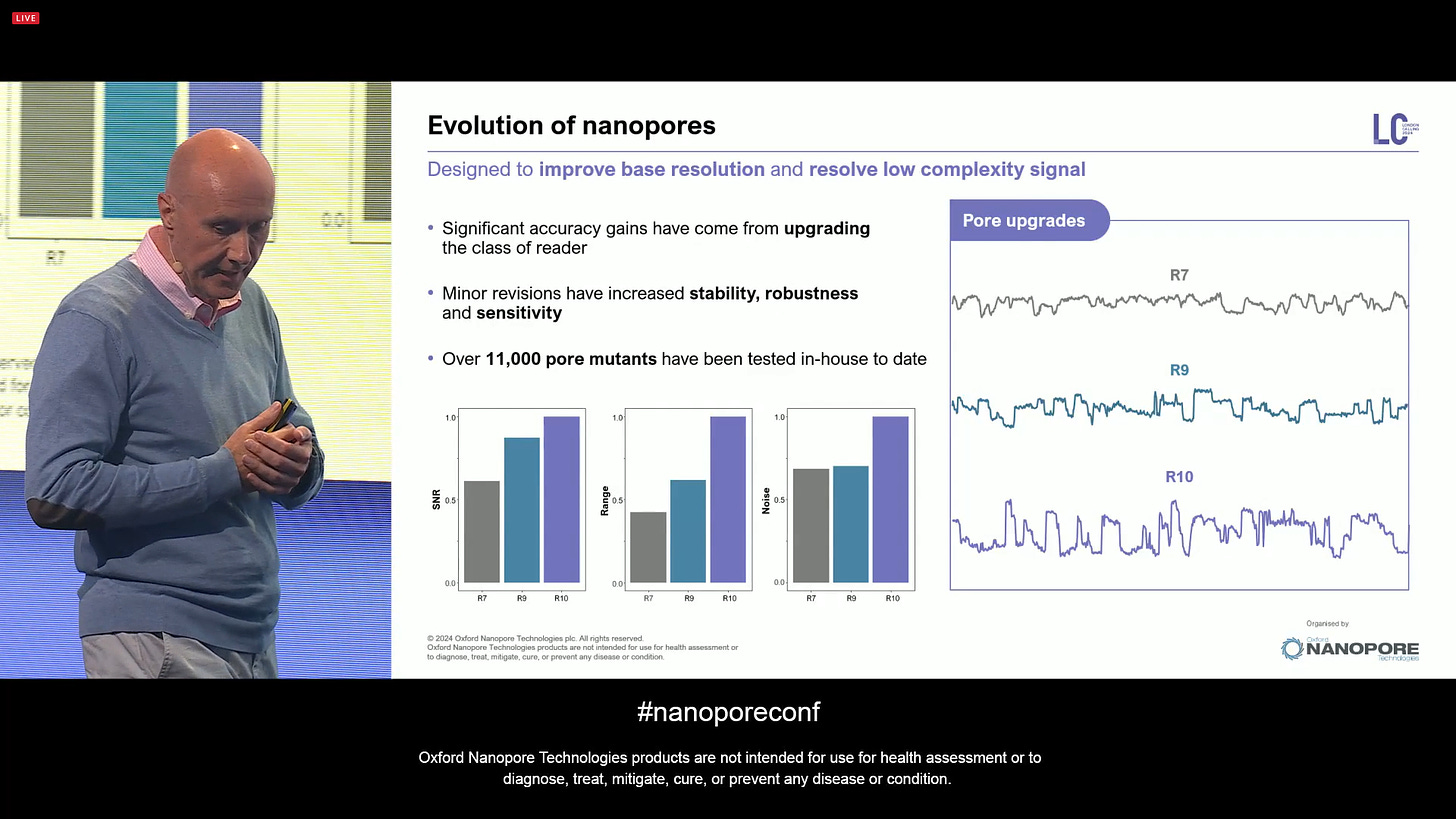
The company went through an evolution of the biological pores themselves, starting with R9, followed by R10, now a pore which has the properties for high accuracy base calling: wide steppy low-noise signal, which traverses the current range frequently. Over 11,000 pore mutants have been tested in-house to date.
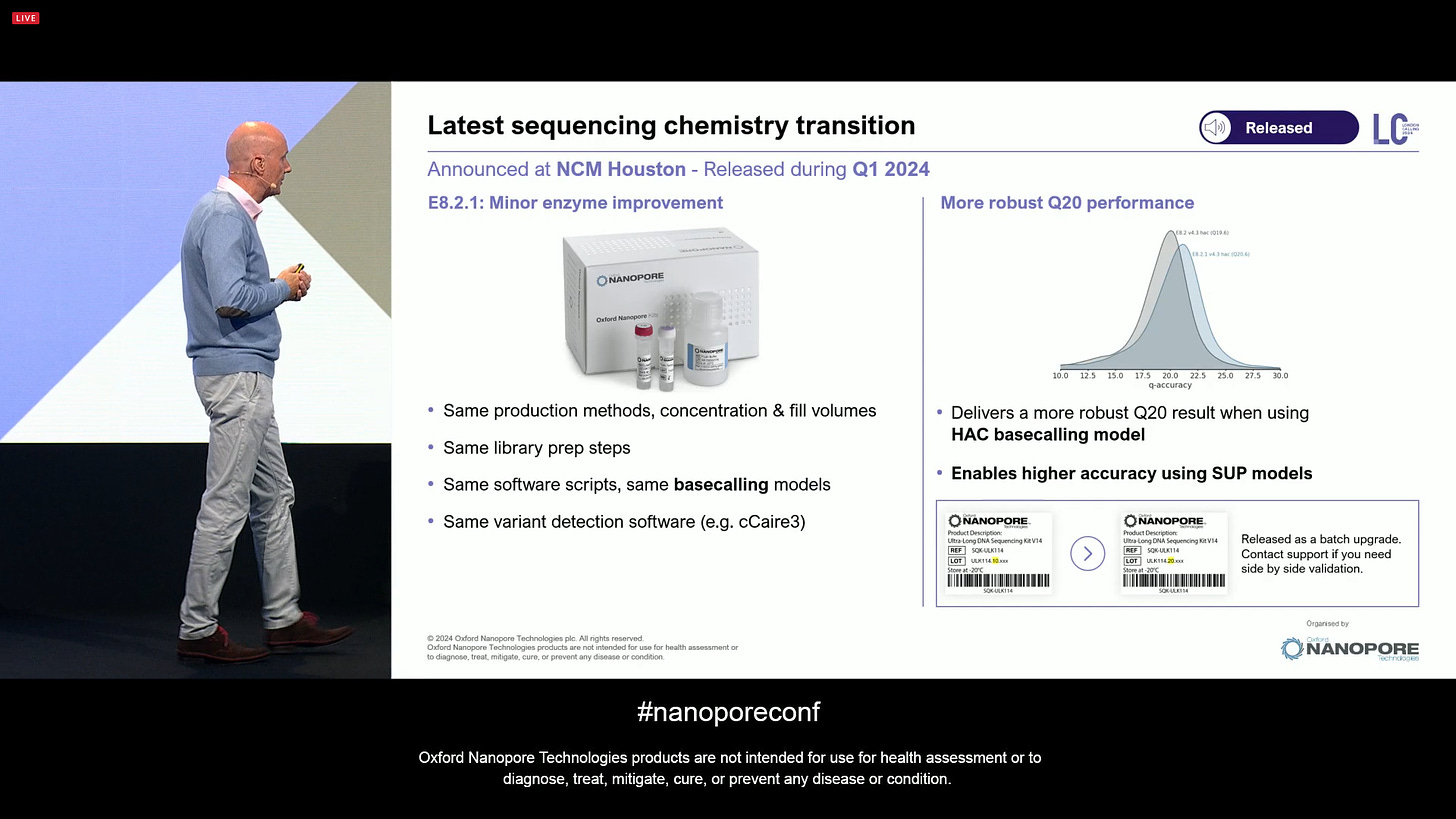
The latest sequencing chemistry transition is quite a small one for the customers: E8.2.1 is a minor enzyme improvement, which has the same production methods, same library prep steps, same software, same basecalling, etc. It gives a more robust Q20 simplex performance, even when using HAC basecalling (SUP not needed). The release is a simple batch update on the product codes.
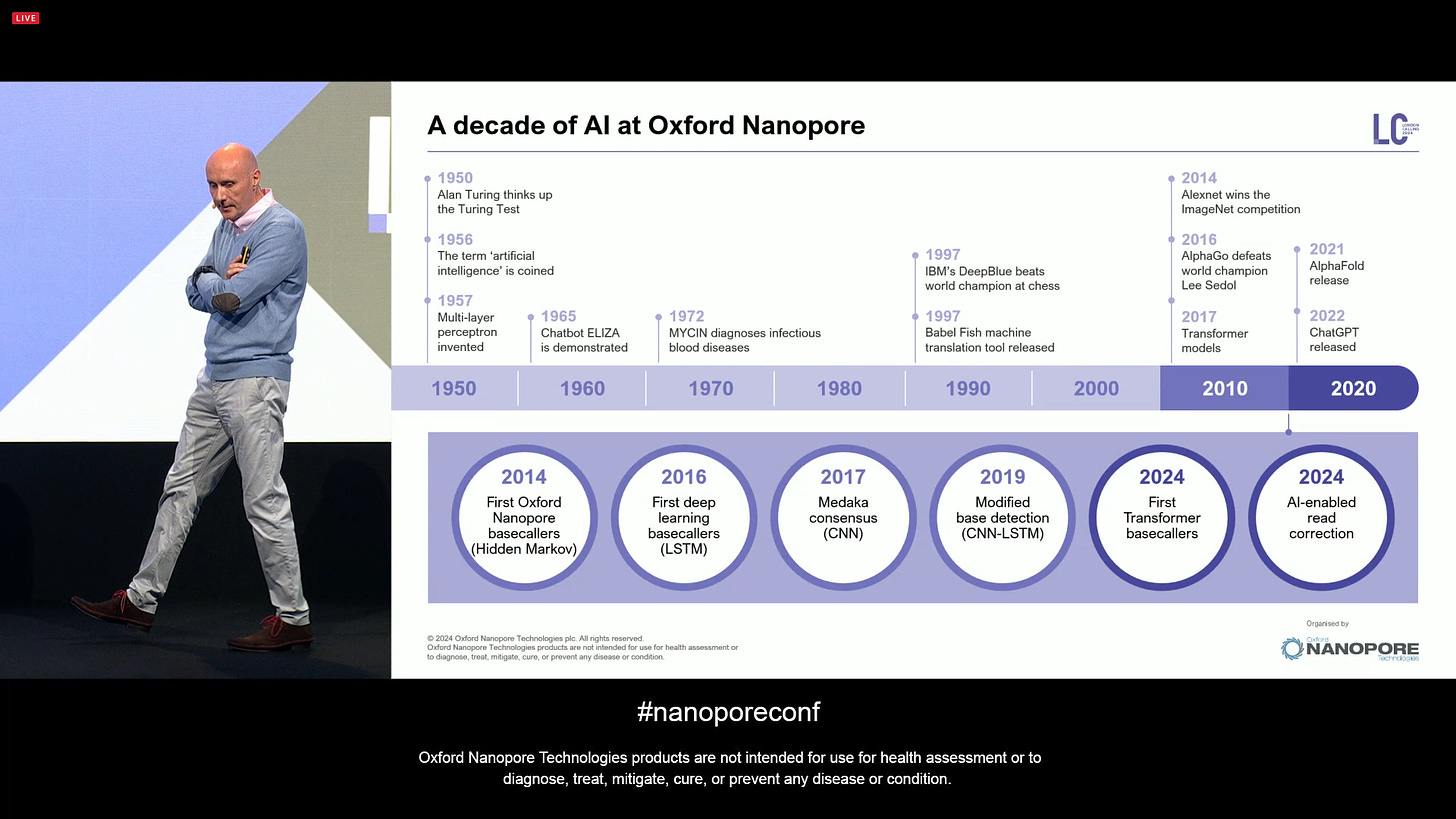
In terms of computation, it’s been already a decade that ONT has been using AI in their products. It all started in the 1950s, when Alan Turing came up with the Turing Test, later on the term ‘Artificial Intelligence’ was coined, and the first Multi-layer perceptron was invented in that decade. At ONT, the first version of the basecallers was based on Hidden Markov Models (HMMs), later on moved on to the first deep learning basecallers with LSTMs, later on using CNNs for Medaka consensus, and CNN-LSTMs for modified base detection, and the big announcement for 2024 is that now the basecallers are based on Transformer models, the same models used in ChatGPT, only here applied to basecalling. The release of the latest dorado also includes AI-enabled read correction (Herro).
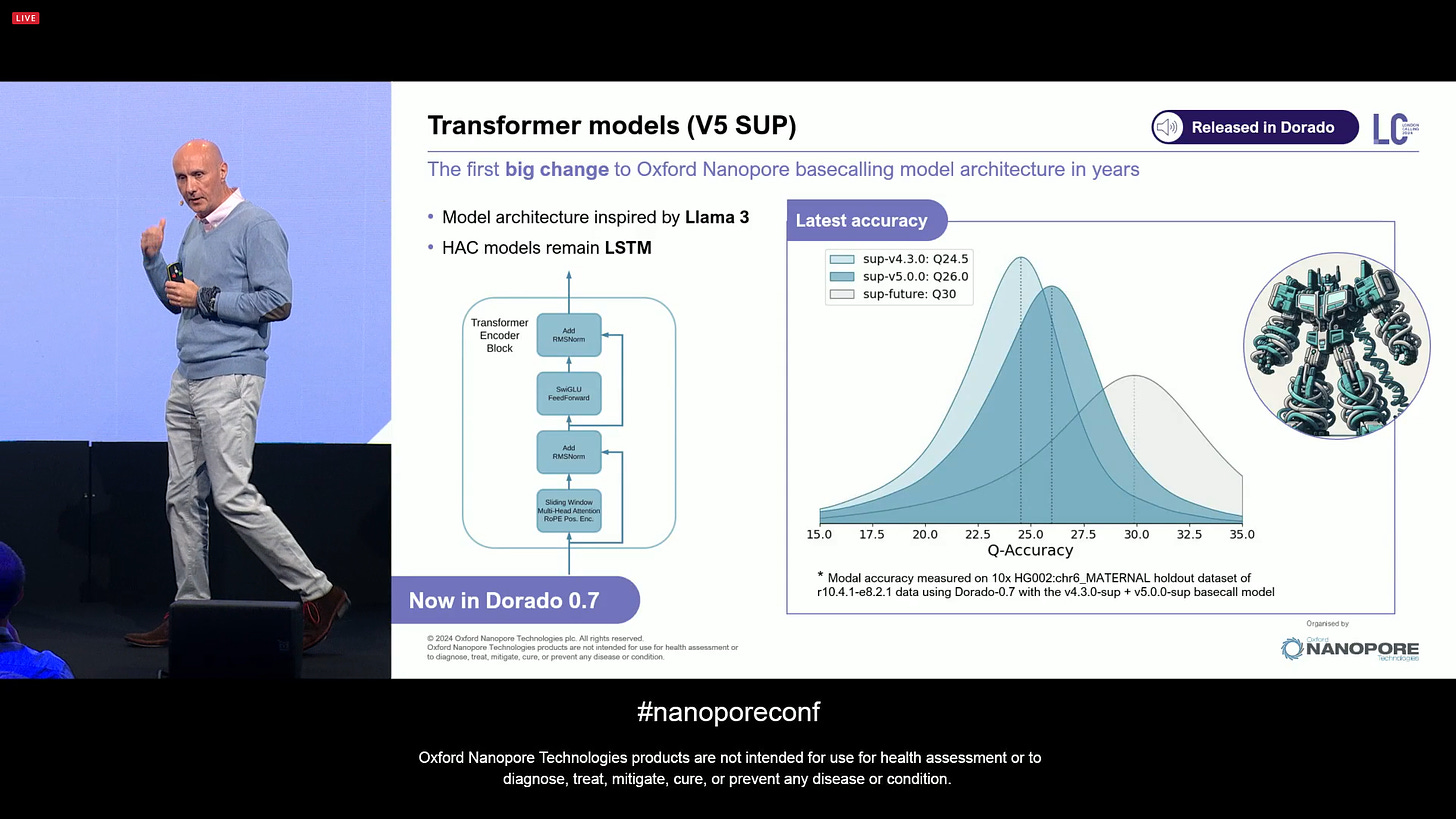
The Transformer models are now released as part of the V5 SUP models. The simplex accuracy jumps from Q24.5 in sup-v4.3.0 to Q26.0 in sup-v5.0.0, and there is an internal version, sup-future, which achieves Q30 simplex data quality (not released yet). The model architecture of this V5 models is inspired by Llama 3 (Facebook Meta), and is now available in Dorado 0.7. The rest of the models remain in V4 LSTM, for example HAC, so that they can still benefit from the performance of years of software tuning. This is with the default 400bps speeds, and is available now. The sup-future is only internal now, as it comes with a -15% yield penalty and 4x performance penalty, and they prefer to keep working on it until it can be released later on. The performance is always a moving target, as GPUs of the likes of Nvidia NVDA 0.00%↑ keep getting faster. Contact ONT if you want to play with the sup-future code on a dev access model.
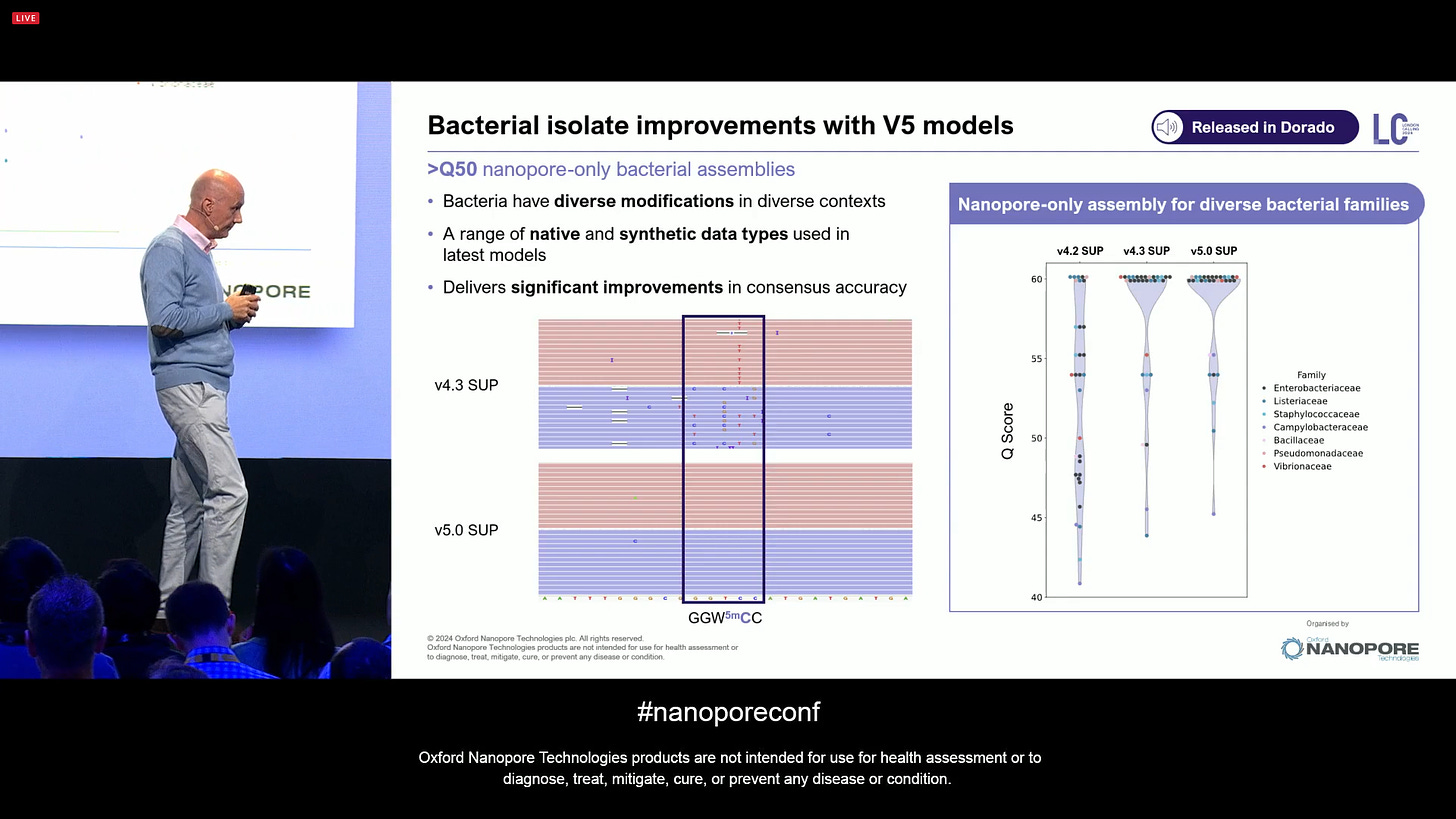
On bacterial genomics, the data quality looks very good with the new V5 models. Part of this is because bacteria/archaea have compact genomes with very few long homopolymers. This means that the ONT technology is getting lots of commercial traction in this segment, as people appreciate the quality of the results that they are getting.
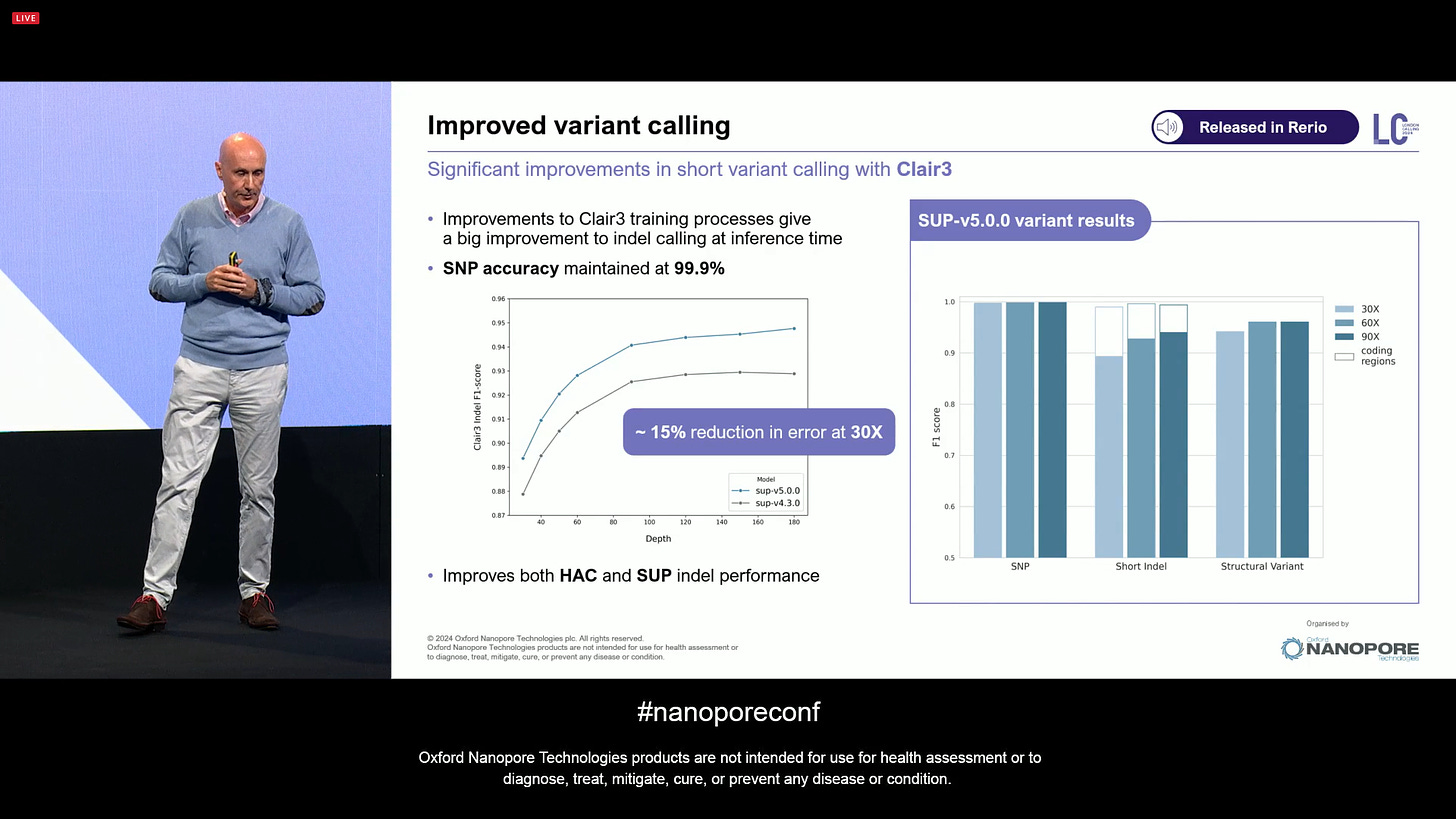
Also improved variant calling, here shown in a collaboration with the developers of the Clair3 variant caller: a reduction in 15% of error at 30X, with the SNP accuracy maintained at 99.9%. The white boxes in the barplots on the right show the extra boost in performance in coding regions. The reason is simple: coding regions rarely have long homopolymers, which demonstrates that the quality of ONT gets an extra boost if measures around coding regions. This is one of the reasons why clinical applications are also getting commercial traction: they care more about coding regions than other parts of the genome.
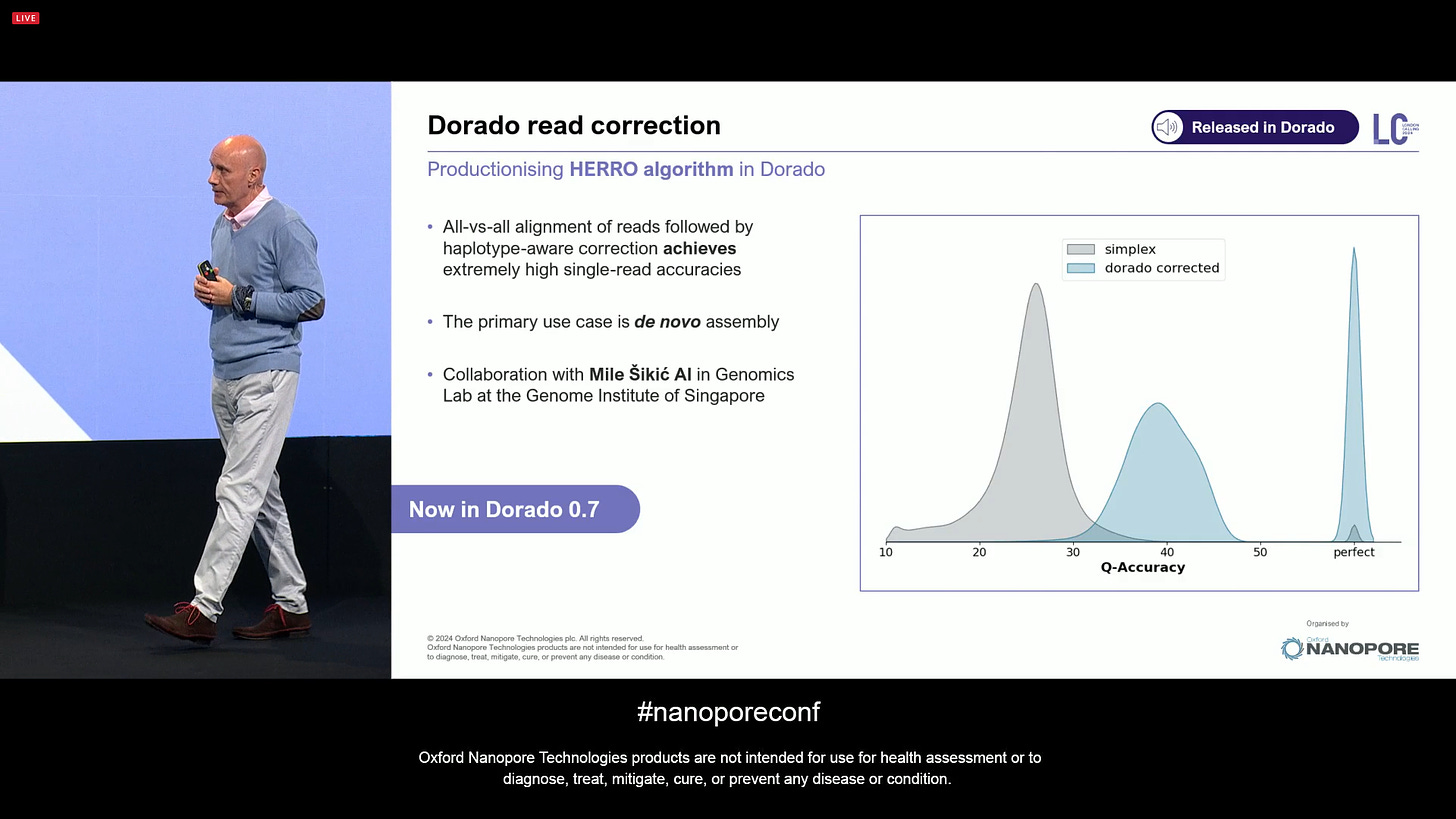
One of the big surprises was the announcement of the AI-based Herro read correction method (there is a biorxiv preprint for the publication of the Mile Sikic AI lab at the Genome Institute in Singapore). In the latest Dorado v0.7, this is now integrated in the ‘dorado correct’ mode. It’s an all-vs-all alignment of reads followed by haplotype-aware corrections, which achieves extremely high single-read accuracy. The primary case is de novo assembly.
One of the main benefits of Herro single-read correction is that it makes the software toolchain a lot easier: people can now take their raw ONT data, basecall it with dorado, then apply ‘dorado correct’ and obtain reads that are very similar in nature to the PacBio HiFi reads that are the input for software tools like Hifiasm or verkko. The later is able to assemble human chromosomes X and Y from telomere-to-telomere (T2T). This if for diploid (multiploid) genomes, but it can also be used with haploid genomes or inbred genomes, in which case using the LJA software gives the best results.
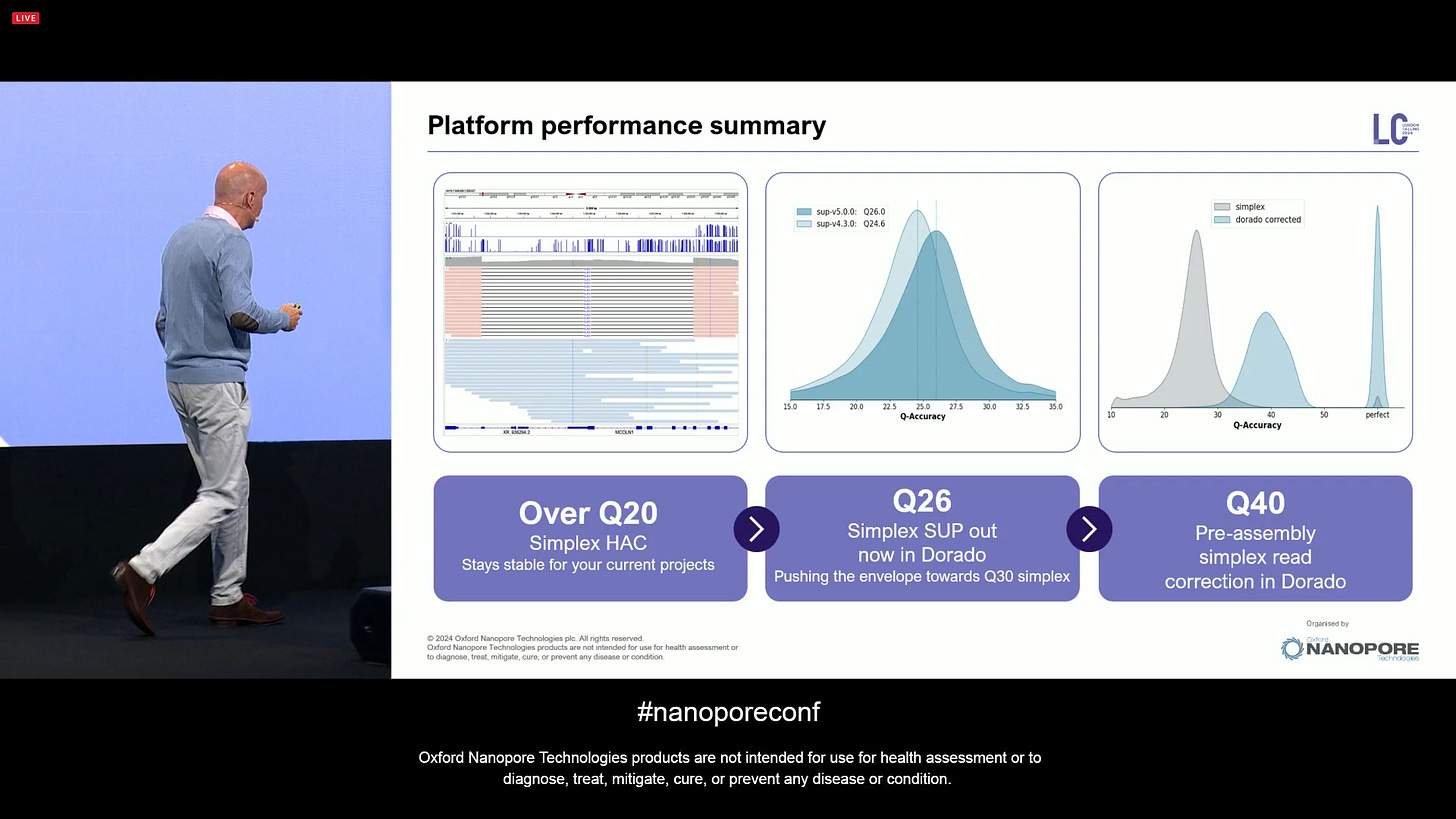
So from Over Q20 with Simplex HAC, to Q26 with Simplex SUP out of Dorado now, one can now also get Q50 pre-assembly simplex reads corrected with ‘dorado correct’ using Herro.
The progress in assembling large mammalian genomes doesn’t stop, what people call telomere-to-telomere assemblies (T2T) has now seen several iterations. A year ago, the best recipe was to get Near-perfect long(ish) reads (20kb) combined with Ultra-long reads (100kb_) and a long range or parental data (Hi-C). This would still have required several technologies.
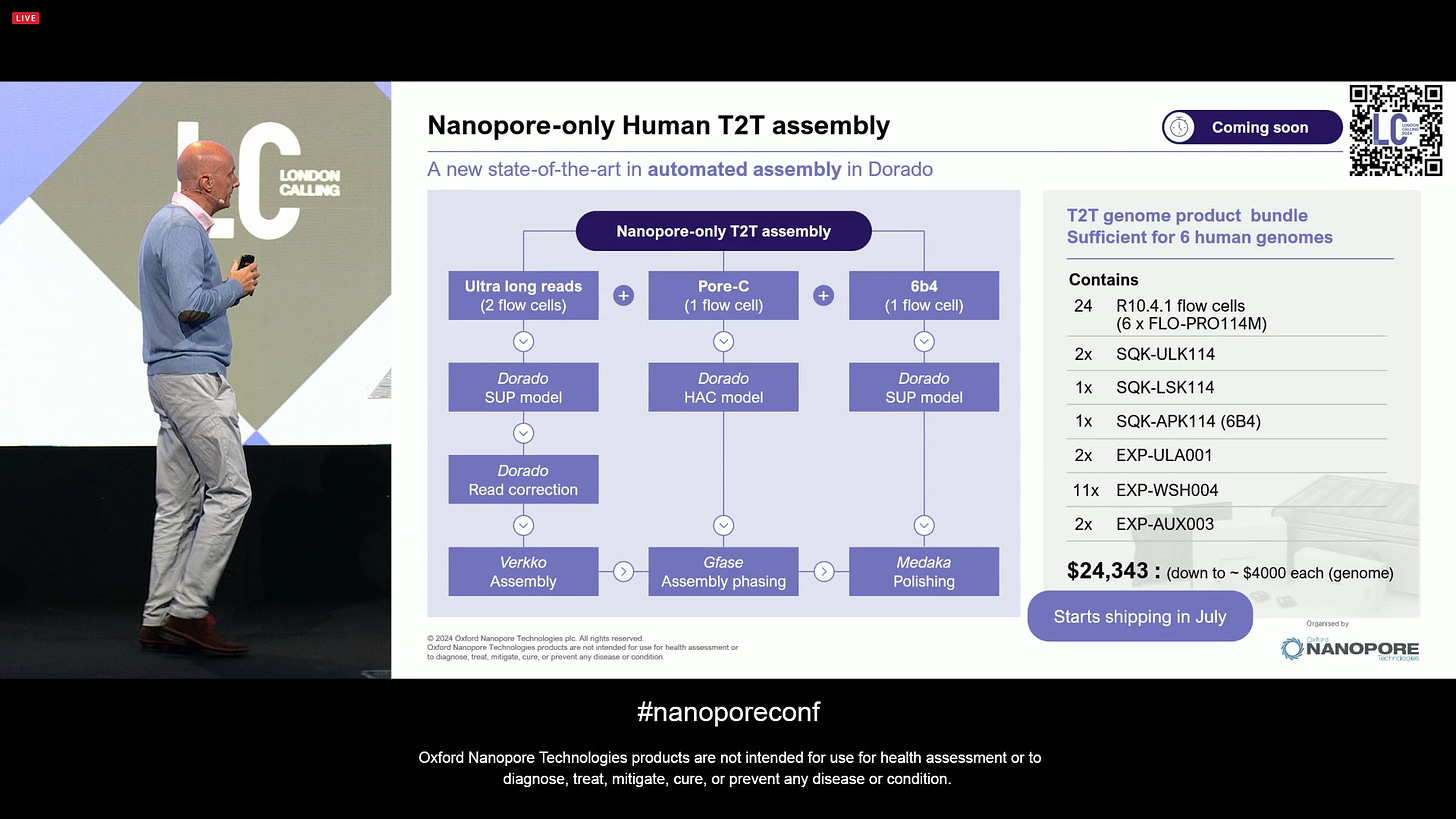
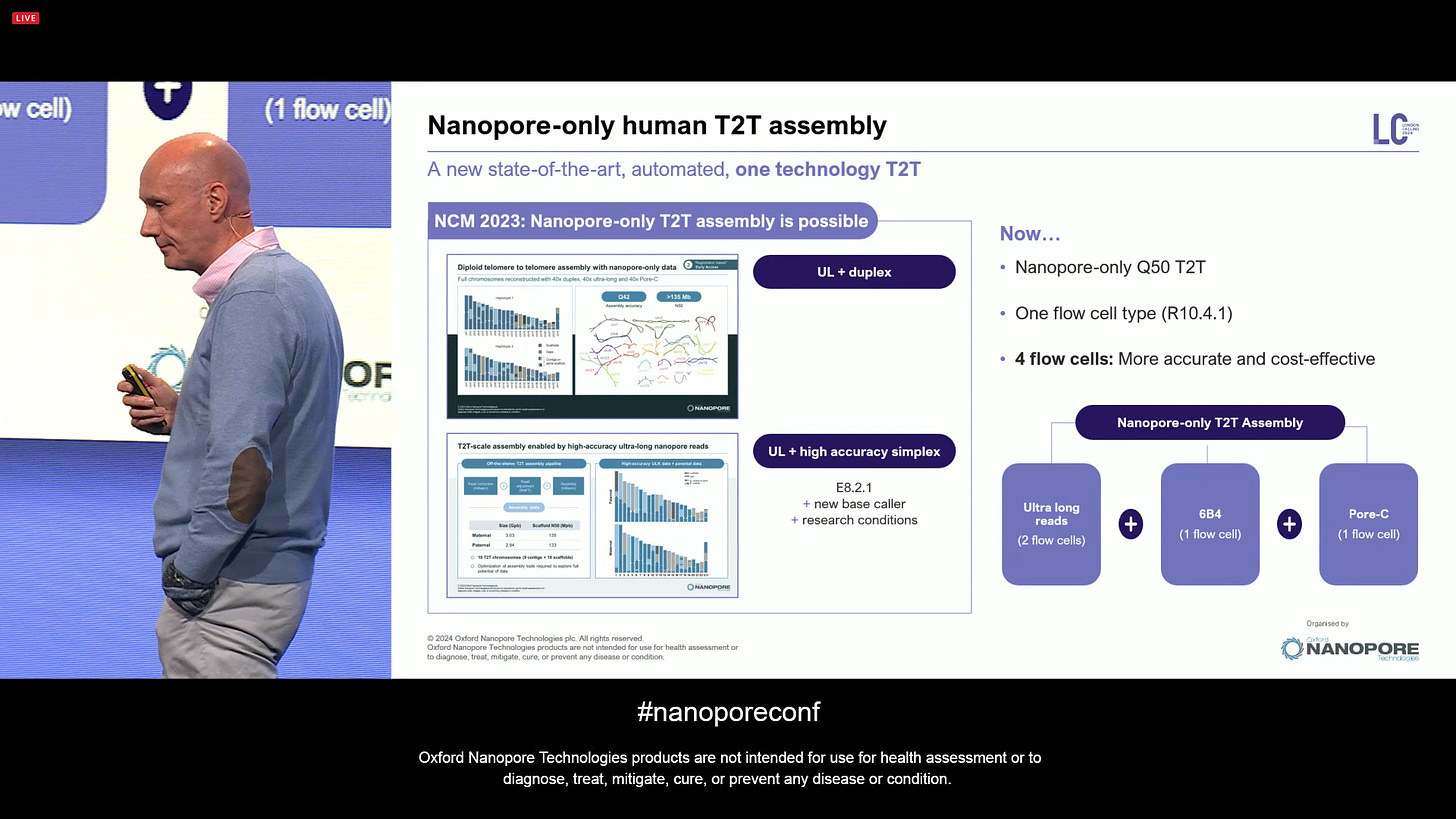
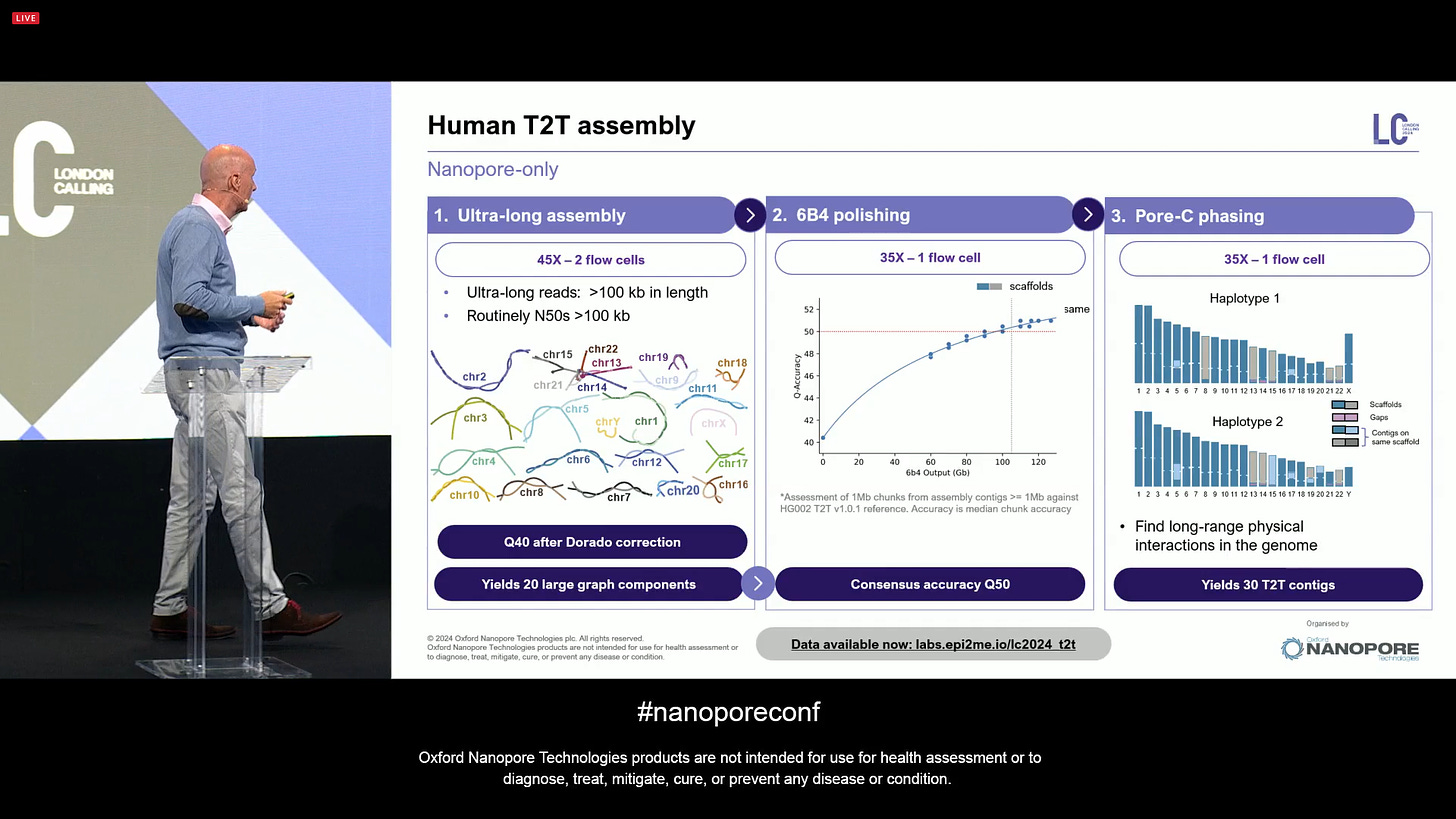
Now aiming at a Nanopore-only human T2T assembly, a new state-of-the-art (SOTA) automated protocol with one technology (it doesn’t need short-reads or PacBio HiFi reads anymore). This involves 2 ULR flowcells, plus one 6B4 flowcell, plus one Pore-C flowcell. It is nanopore-only, one flow cell type, all producing human T2T data with only 4 total ONT flowcells.
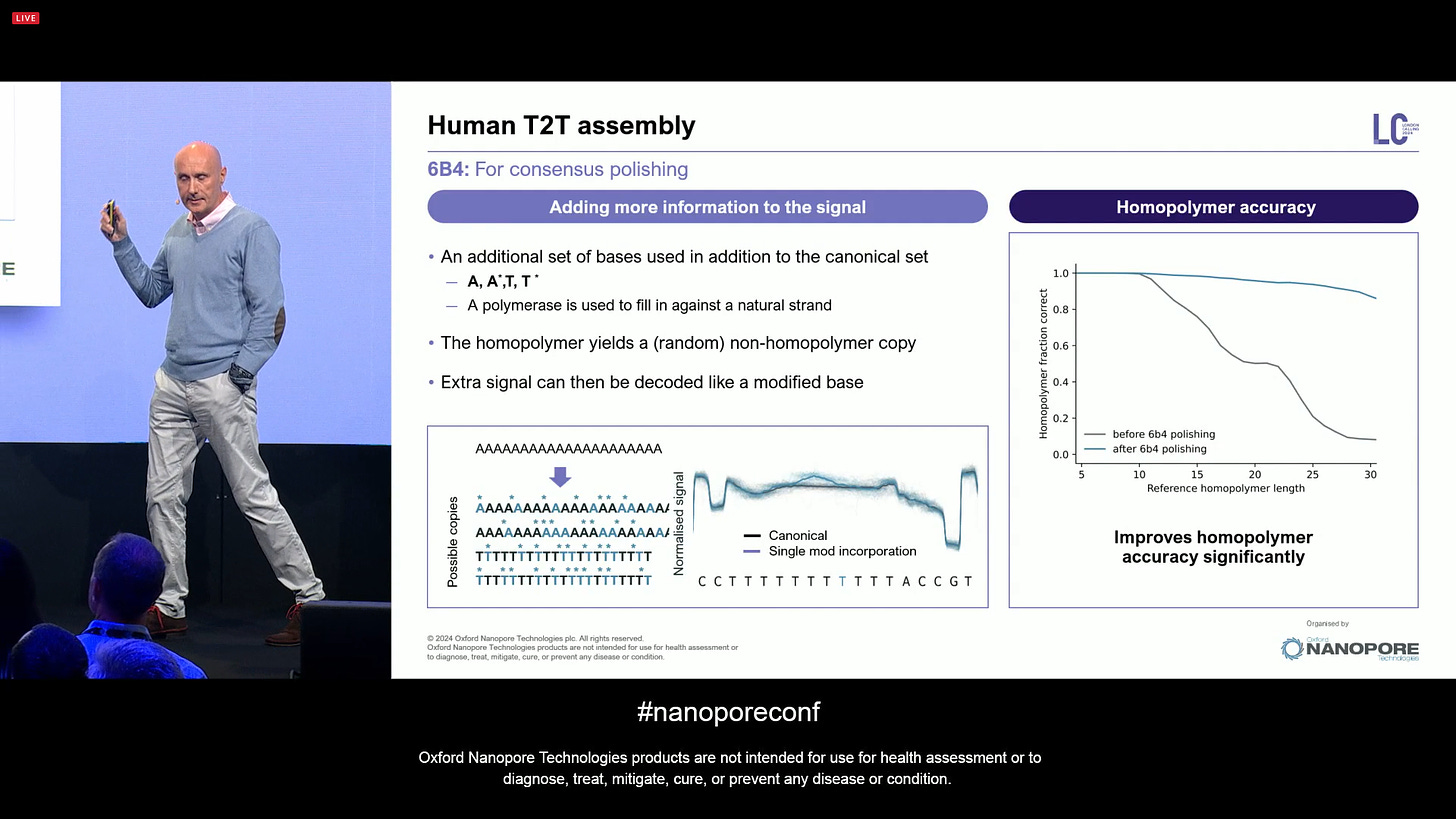
Starting with the 6B4 part: this is for consensus polishing, concentrating on long homopolymers. It is based on the addition of a set of modified based to the canonical set during a PCR reaction. The 6B4 is an upgraded version from the 5B4 protocol that was presented last year: A,A*,T,T* mixed together, which de-flattens the signal, giving a better estimate of homopolymers. This is now coming out as a product. It largely resolves the consensus quality in short indels in homopolymers.
Now the human T2T assembly combining these 3 preps with ONT-only sequencing is able to assemble 30 T2T contigs (out of the total of 23x2 in a human diploid genome).

This is all now bundled up into a kit that will be sufficient to sequence and T2T assemble 6 human genomes, all included, for $4,000 each T2T genome.
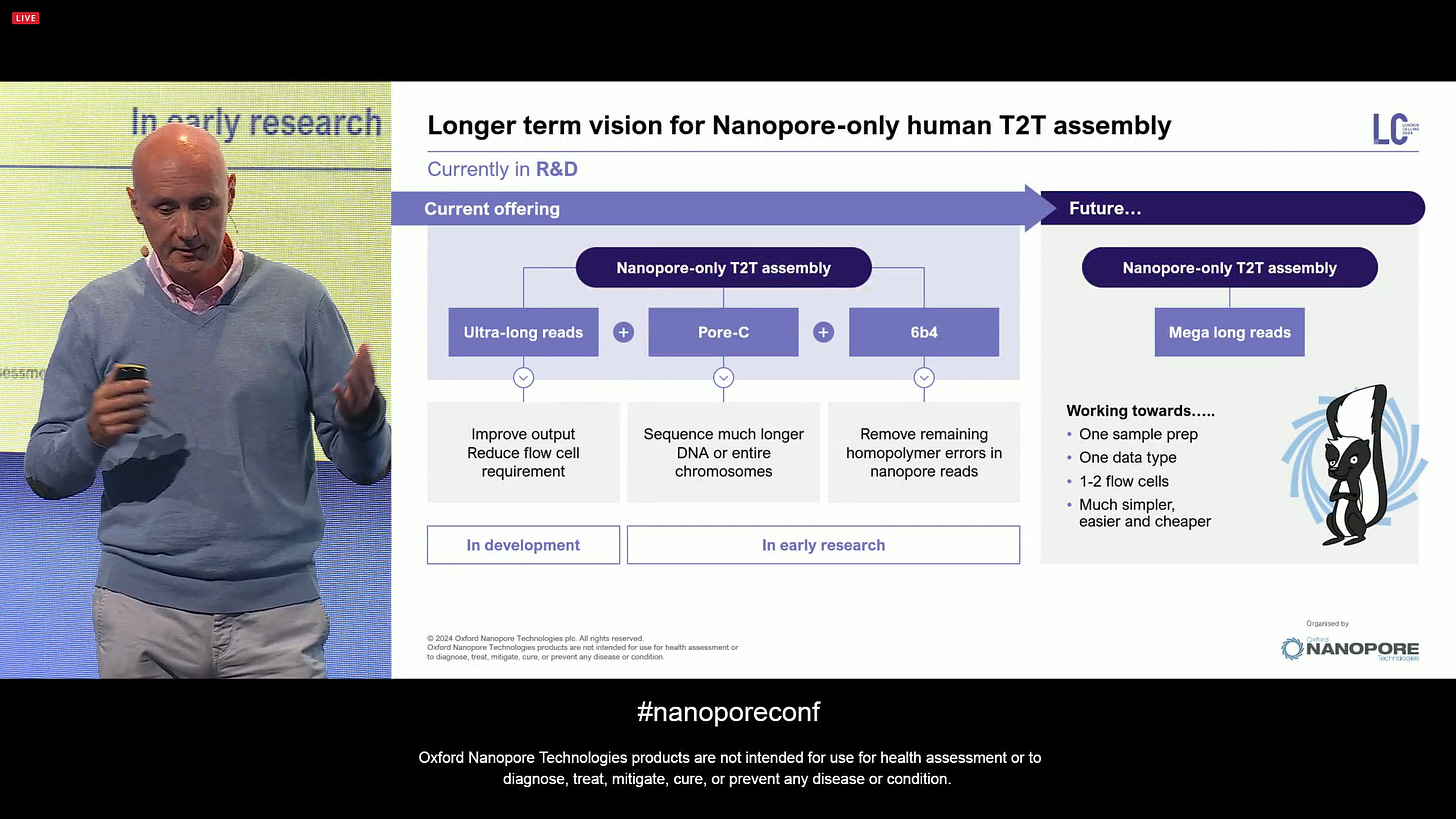
But the longer term vision for Nanopore-only human T2T assembly is even more ambitious: one sample prep, one data type, 1-2 flow cells, and a much simple, easier and cheaper overall experience.
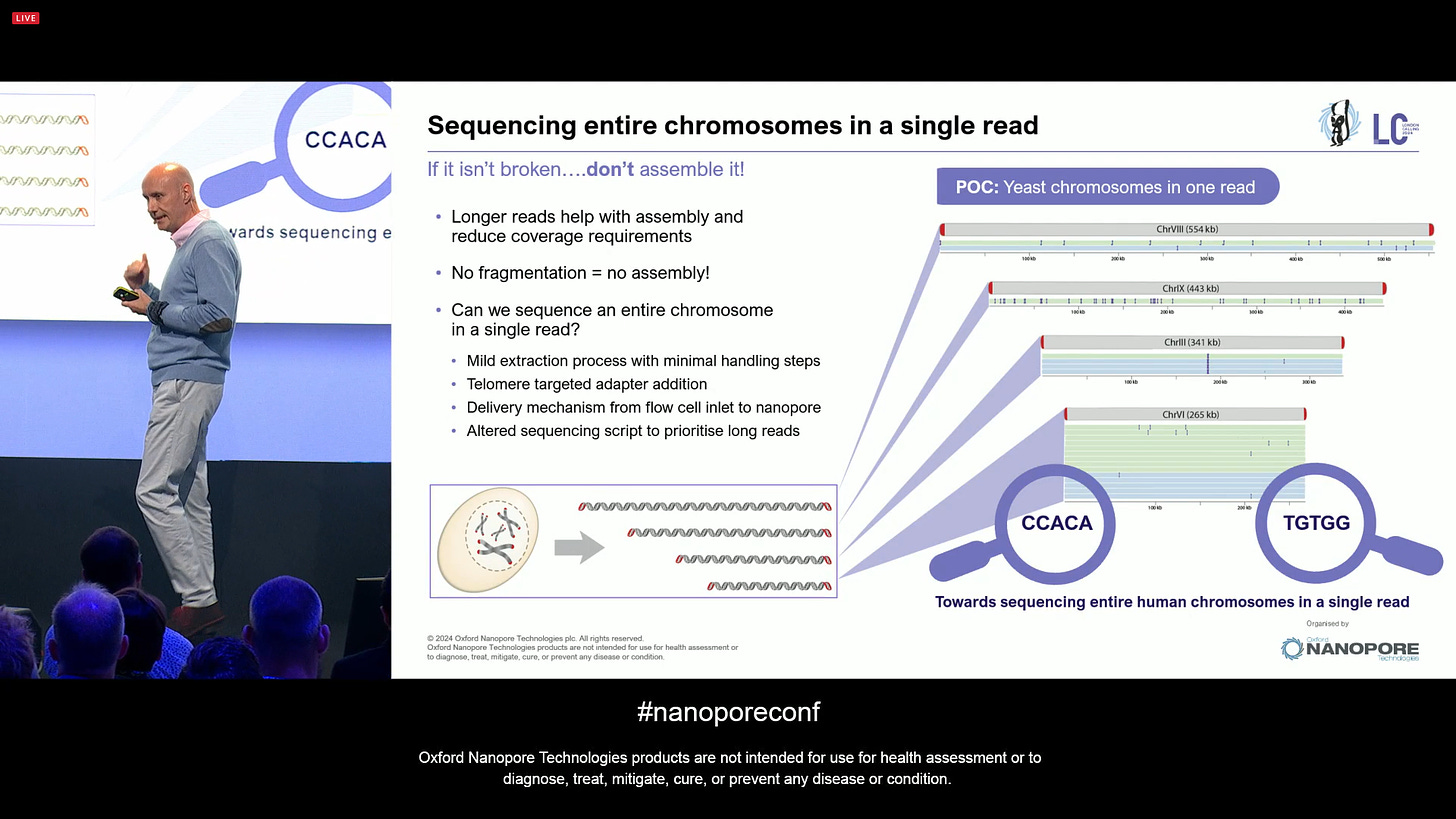
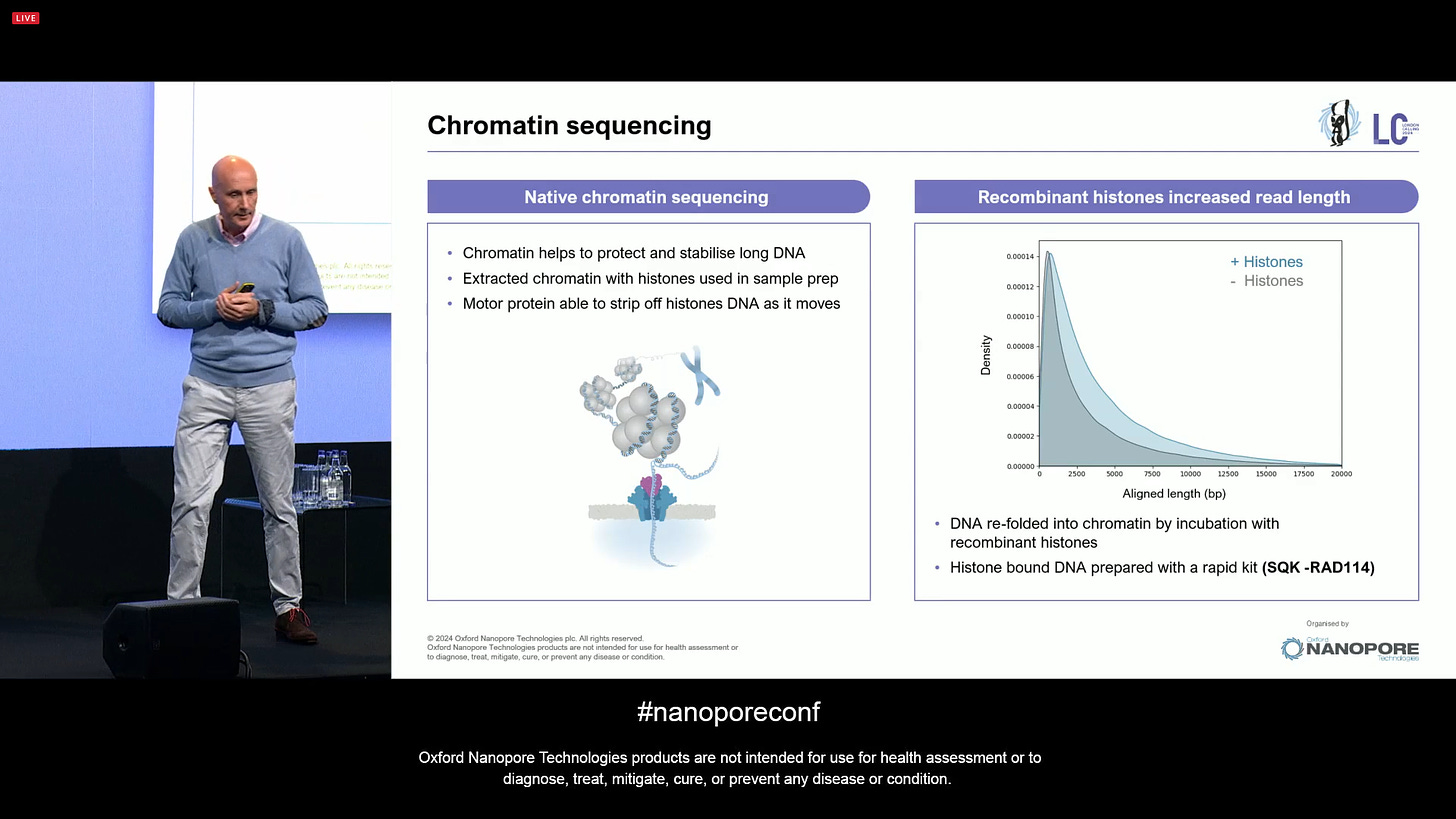
In order to get to that point, one approach could be sequencing entire chromosomes with a single read. If it isn’t broken… don’t assemble it! In order to do this, one needs very unconventional extraction methods, but these exist since the 1970s, and they are perfectly feasible to apply here. Given other tweaks, like designing adaptors that only target telomeres, then physically transporting this to the flowcell without having to put it through narrow gauge tips, then making sure the pores survive when sequencing entire chromosome strands, all goes in the direction of going from “ultra long reads” to “mega long reads”. ONT has been trying this with yeast chromosomes, as is now optimizing it for human-sized chromosomes. If this could happen for single cells, it could transform the way people do biology in many ways.
Some learning from this experiment is that the nanopore sequencing method can deal with chromatin, the proteins that help protect and stabilise long DNA. Since the motor protein that ONT uses is able to strip off the histones as the DNA moves through the pore, it makes the chromosome-level sequencing feasible even without having to worry about chromatin.

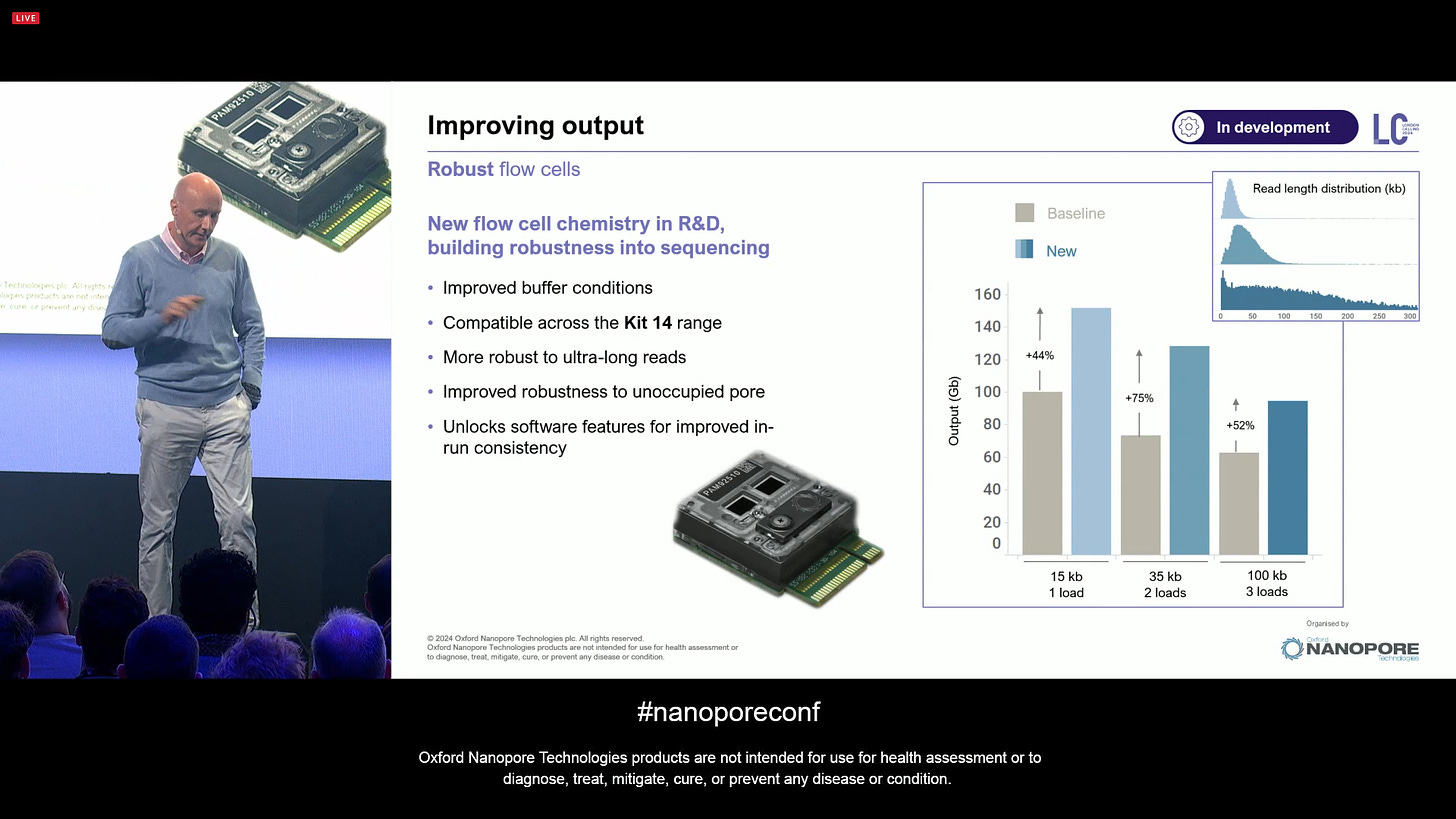
More advancements in flow cell design: ONT has made steps to make flow cells more robust, improving the buffer conditions, which is a small but significant change that can be rolled out transparently. This means the unoccupied pores are now surviving for longer, and it unlock software features for improved in-run consistency. This will raise yields in gains that are equivalent between flushing and reloading a run every 24 hours.

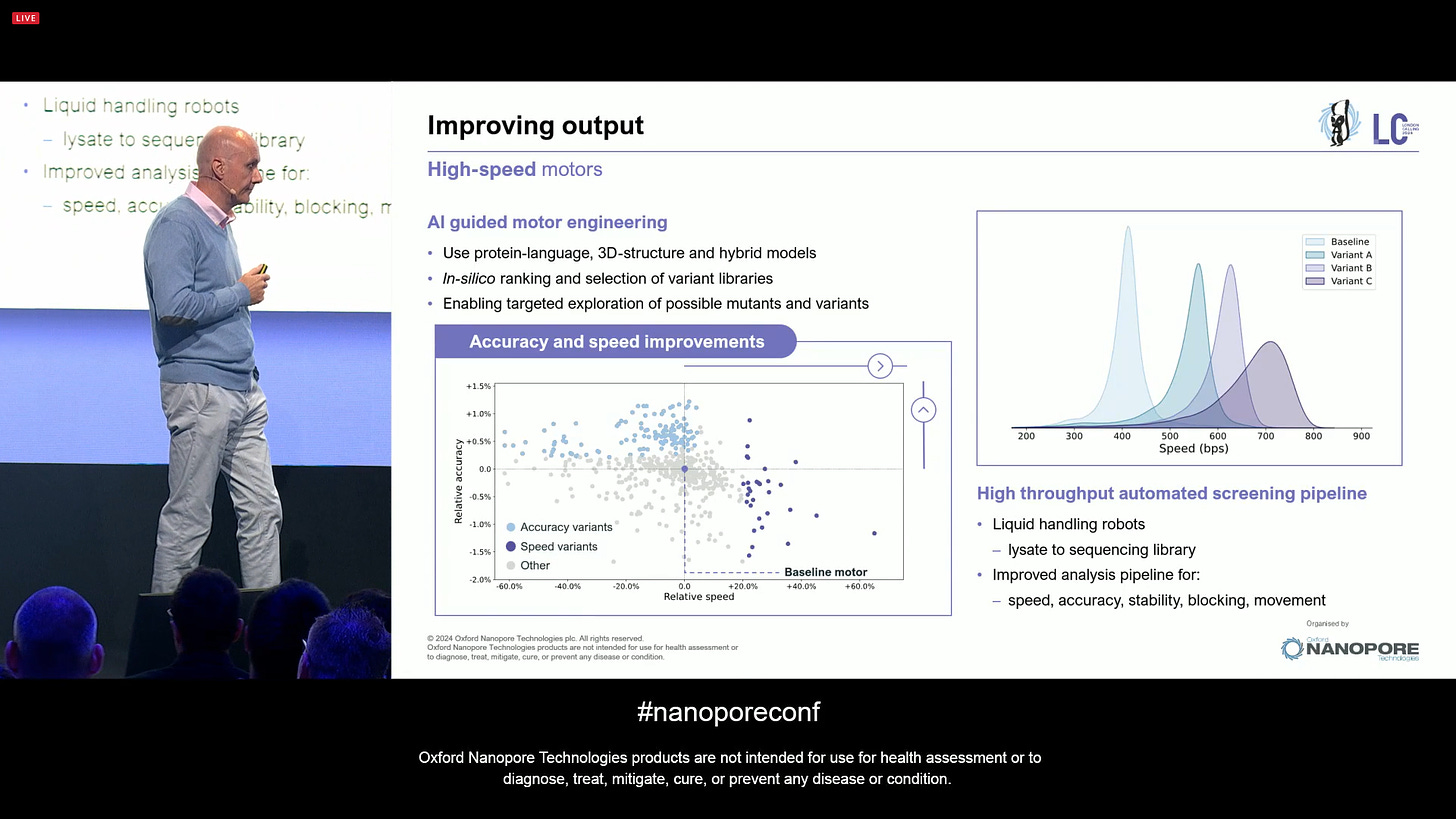
ther step towards improving the output is the work on High-speed motors. ONT has been doing a lot of work that is guided by Artificial Intelligence: using protein-language and 3D-structure models of these enzymatic motors, in-silico ranking them to select from variant libraries, it enables a targeted exploration of possible mutants and variants. ONT can now internally run at much faster speeds while maintaining accuracy, which is another step towards higher yields.
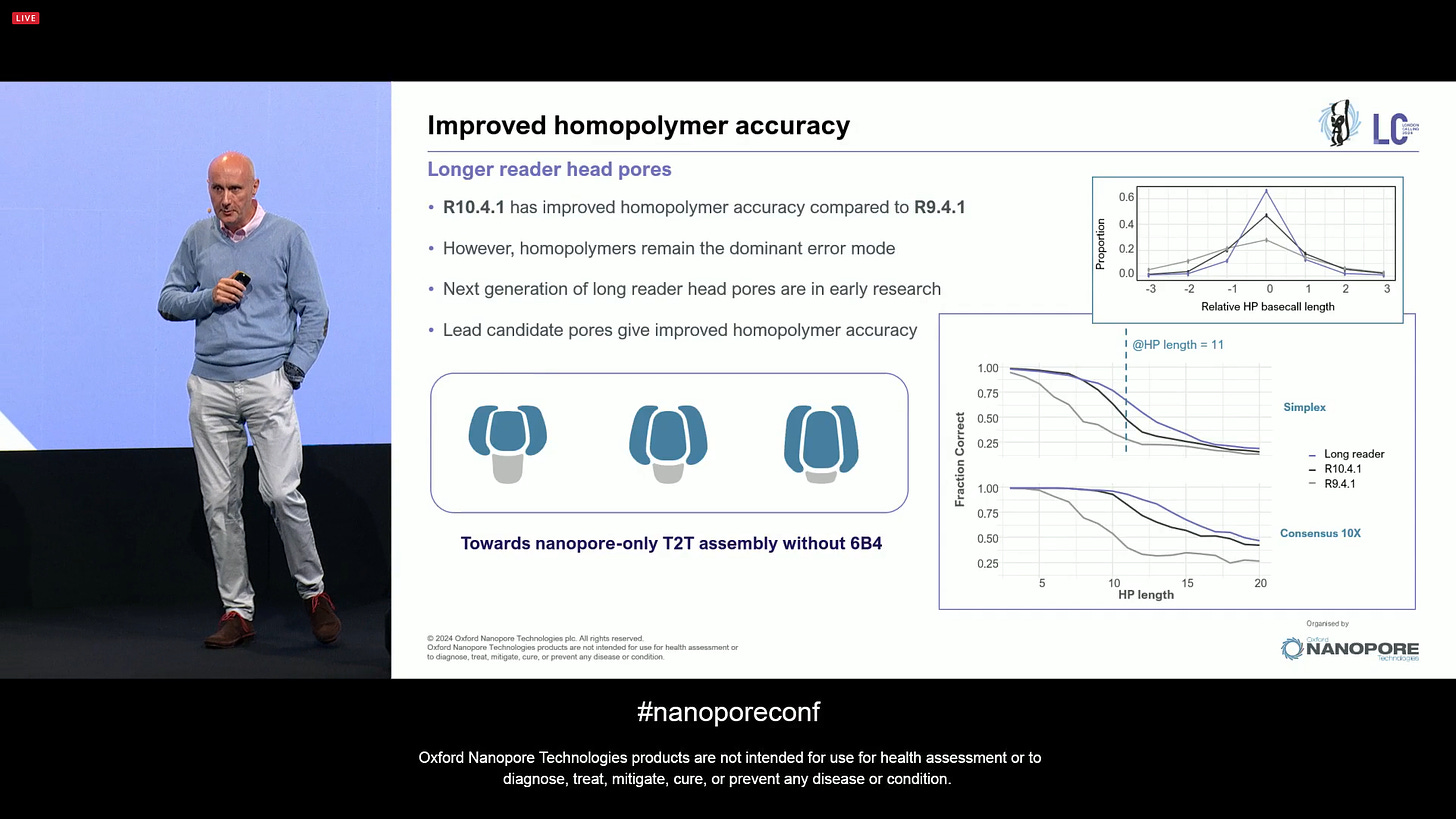
ONT has also been working on improves homopolymer accuracy, and continues to investigate new pore designs. A question here is “how long can a biological pore be?” and the company has already been challenging opinions in the field. Some of the new pore designs show better homopolymers, all adding up to higher accuracy across all contexts.
Moving on to modifications, this is an area that is still difficult to grasp for many new entrants to NGS and genomics (and epigenomics). The current tools are sometimes so bad that limit what people can imagine they could do in their research, and ONT is challenging the current perception of the limits of epigenetic profiling.
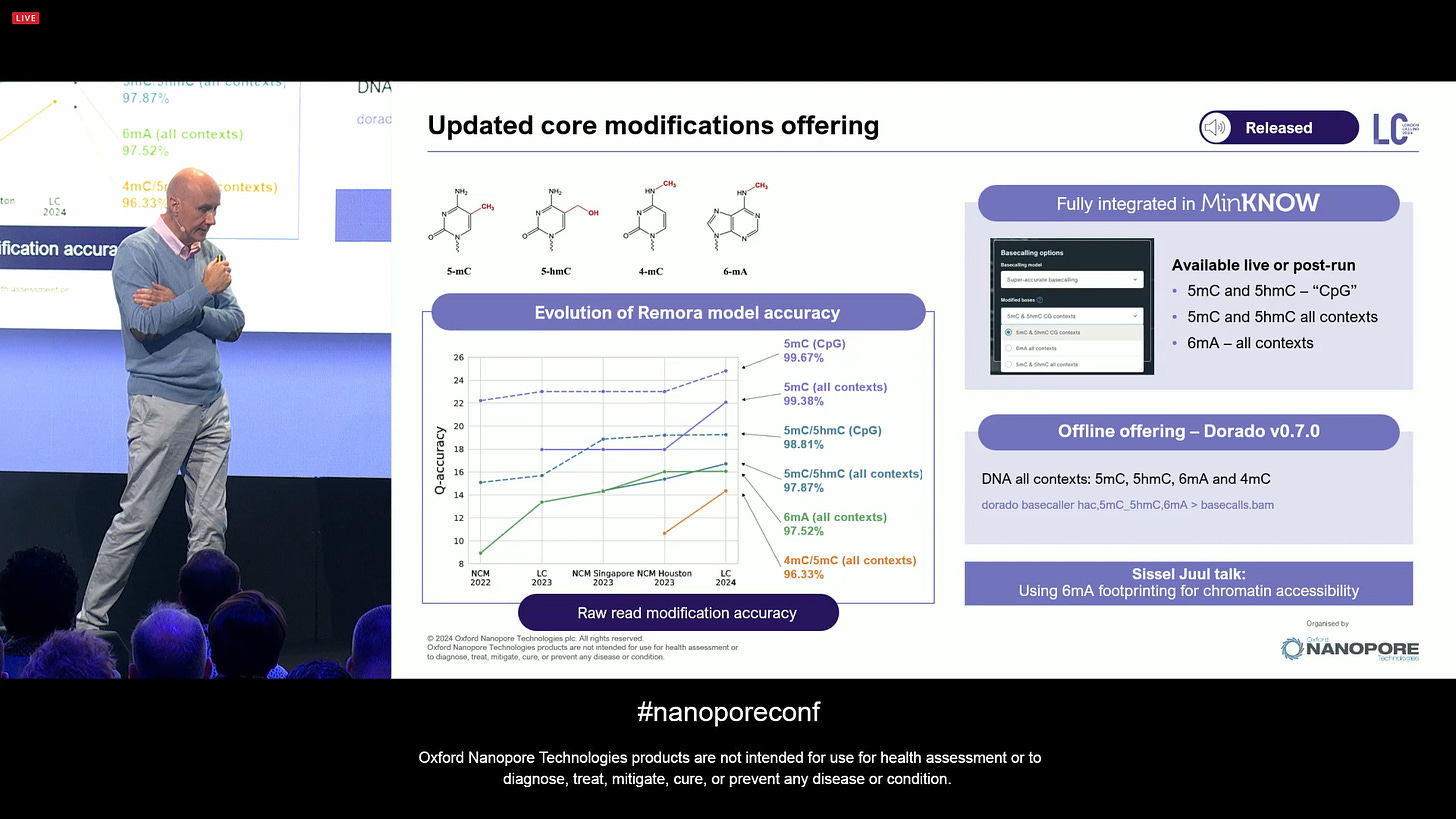
There has been ongoing effort to improve the accuracy of modification basecalling (mod calling). This is for some of the most useful mods in human genomics, like 5mC and 5hmC, but extends to any kind of DNA mod. The list of supported mods keeps growing, and ONT is continuing to tackle more and more of them.
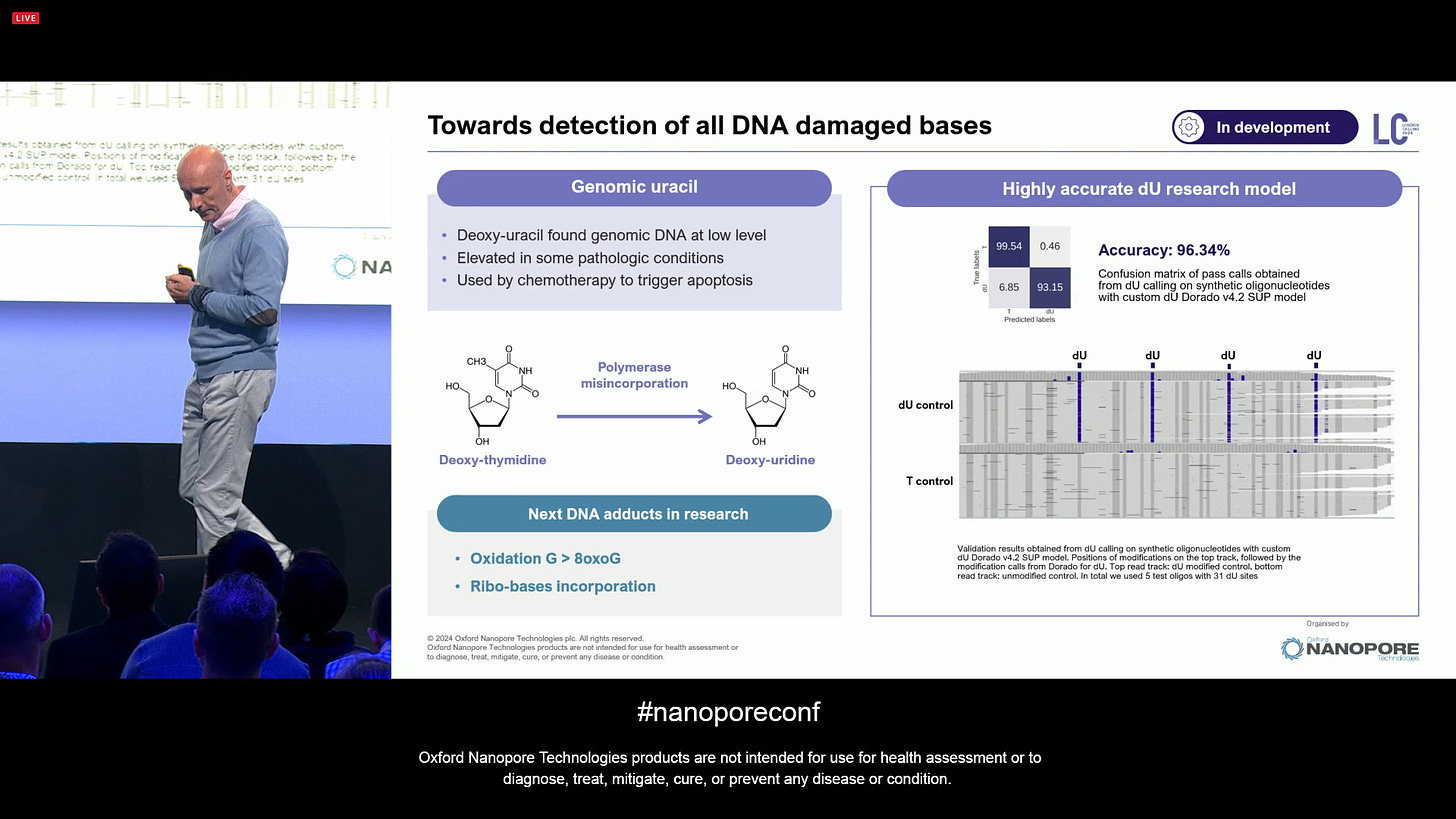
ONT has continued to work on DNA damaged bases, such as deoxy-thymidine and deoxy-uridine. There are more in the list, including 8oxoG and ribo-bases. This will undoubtedly lead to a better understanding of what causes these damages, and what information can be extracted from it that is relevant, e.g. for human health. What happens when someone received chemotherapy or radiotherapy? Which bases get damaged?
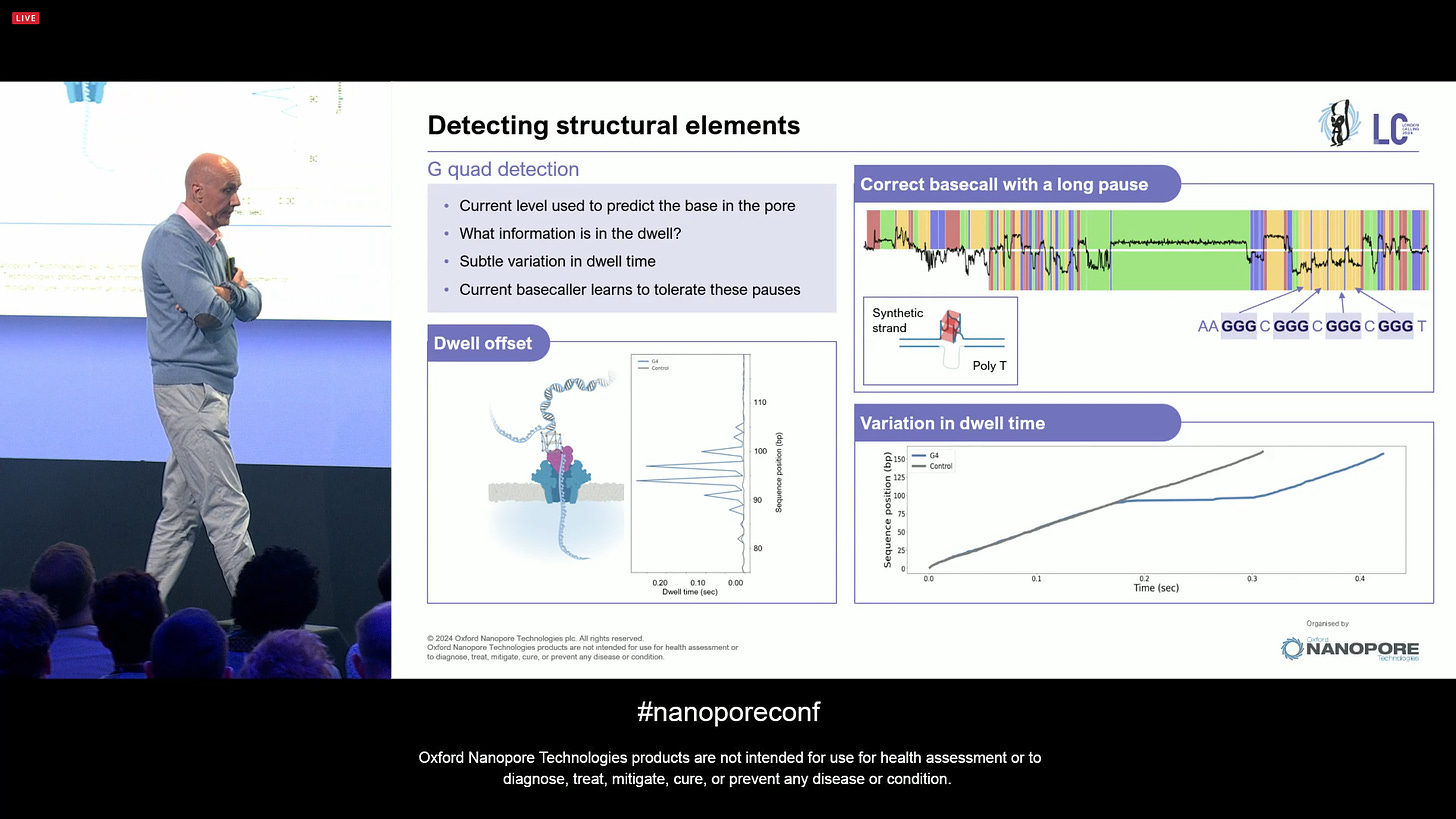
A slight diversion here is the detection of structural conformations of DNA, including G-quad detection. A lot of this is driven by software.
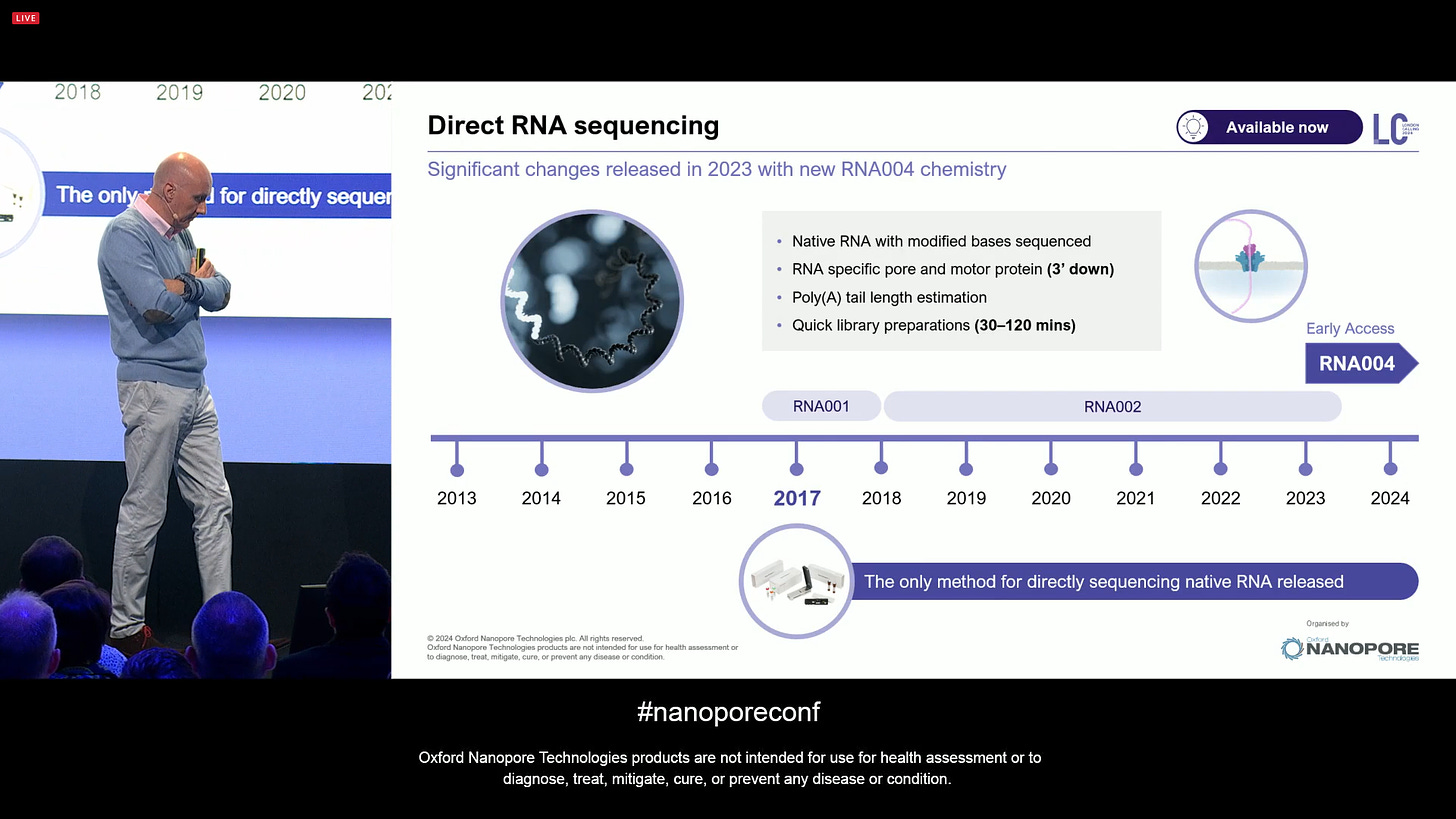
Moving on, it’s now time to update on Direct RNA sequencing. This was an idea that came from the Applications group, so in many ways driven by what kind of Science would one be able to do if the technology was there to perform these type of assay. As a product, it has had consistent growth in interest and uptake, and it keeps getting bigger.
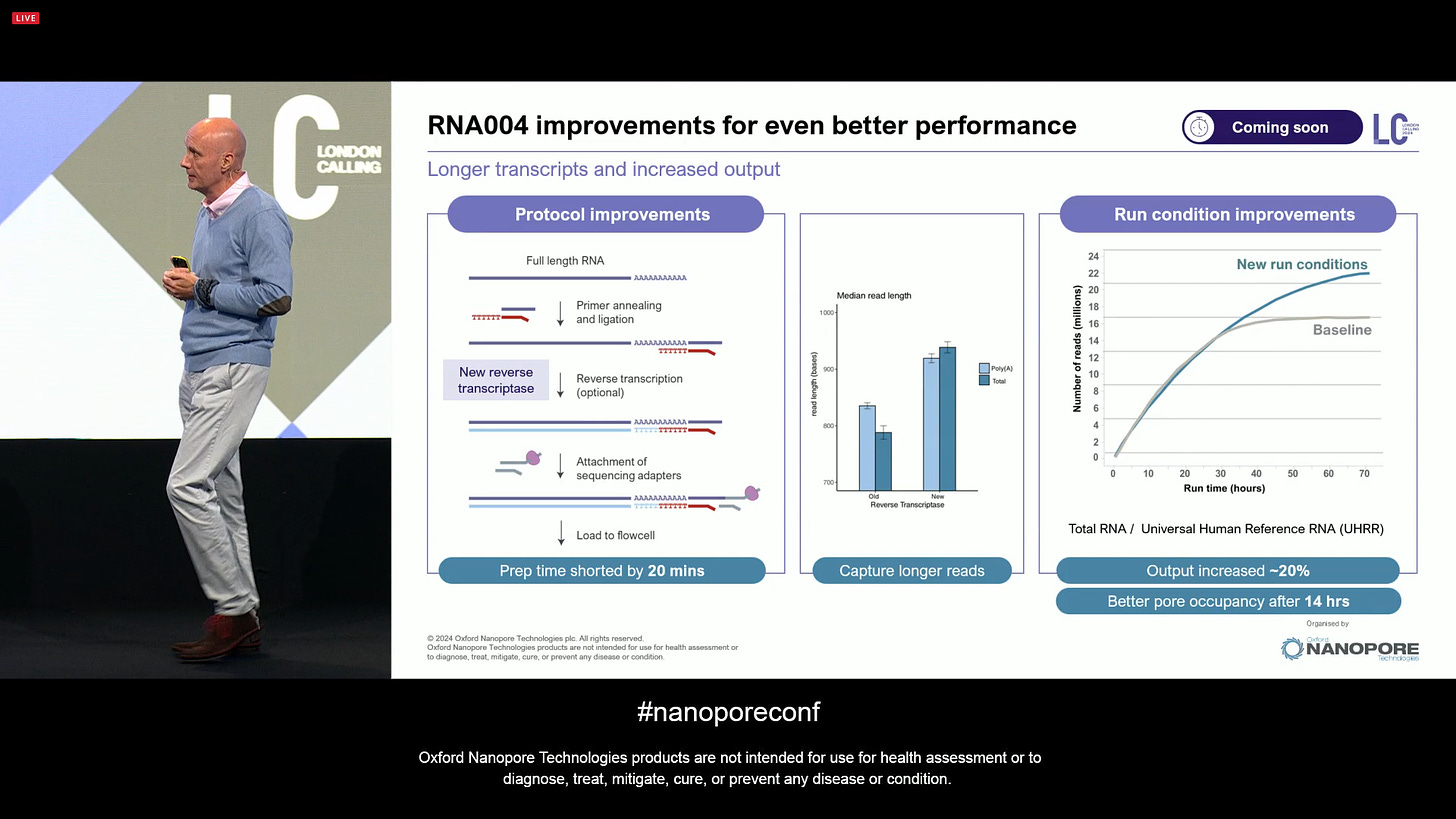
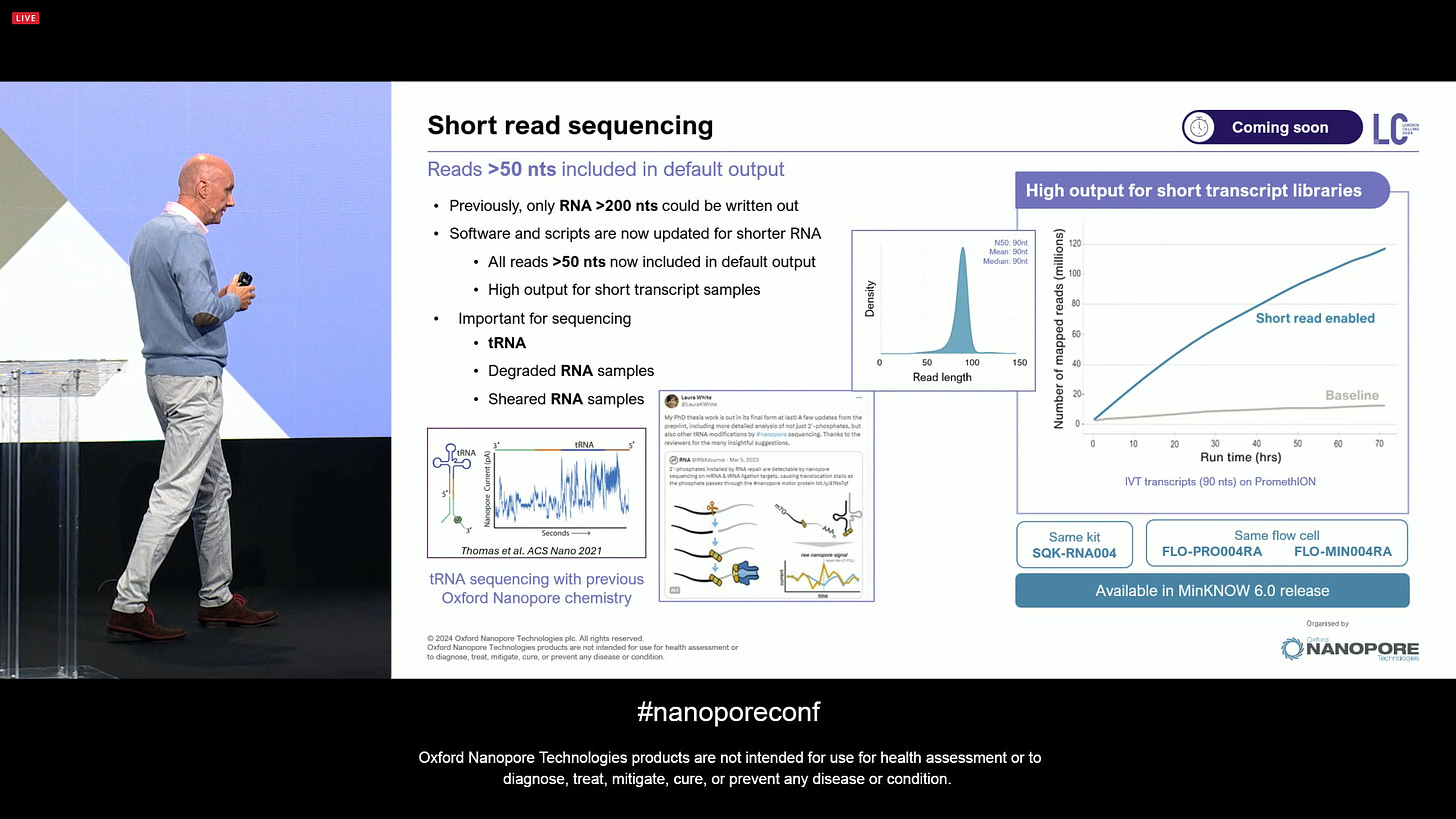
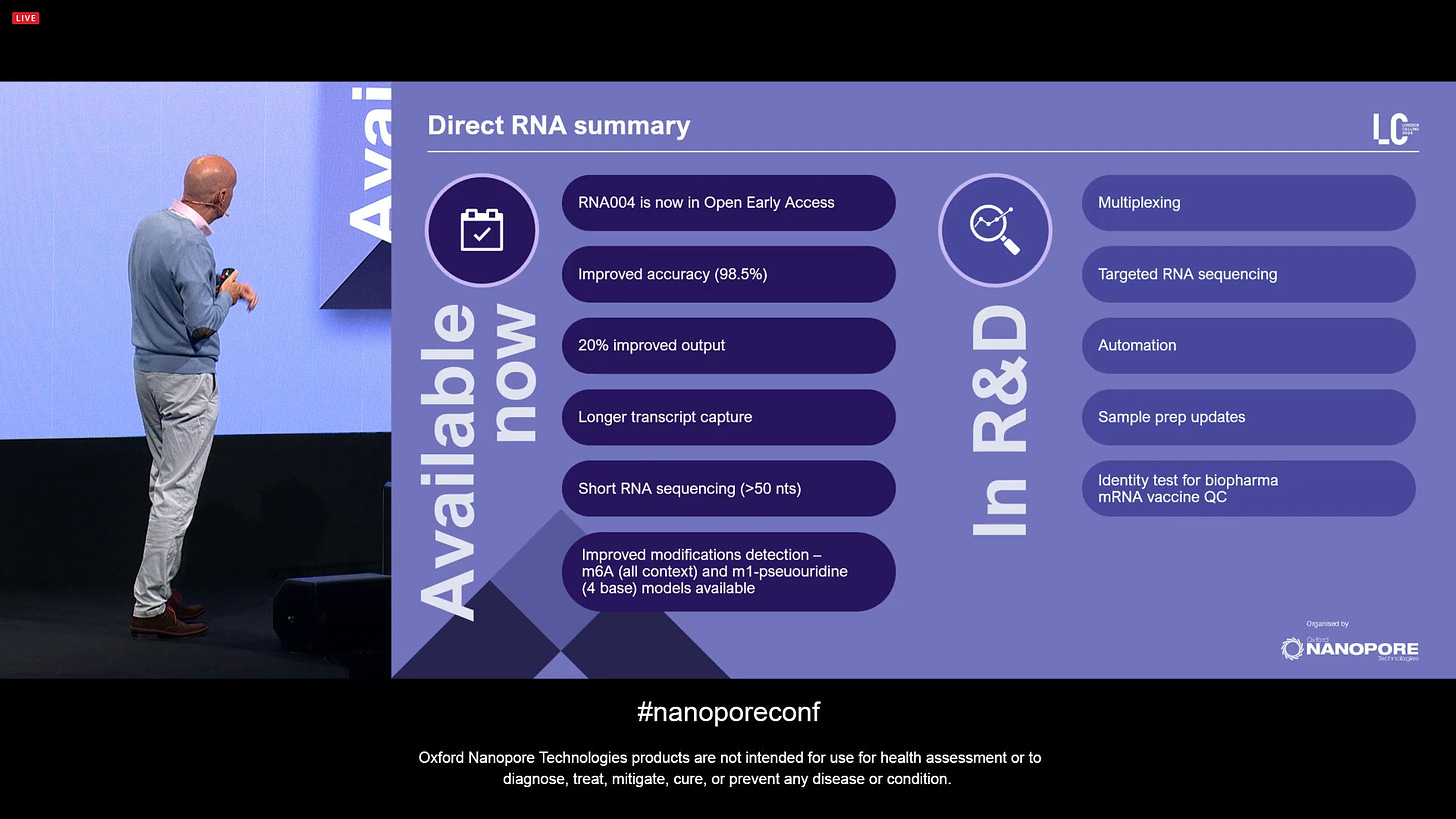
ONT announced further improvements to Direct RNA sequencing for the RNA004 version, with longer transcripts and increased output.
Also a software update that means the Direct RNA method is now able to detect short reads, going from a minimum of 200 nts to now 50 nts included by default. This includes improtant types of RNA, such as tRNAs, degraded or sheared samples.
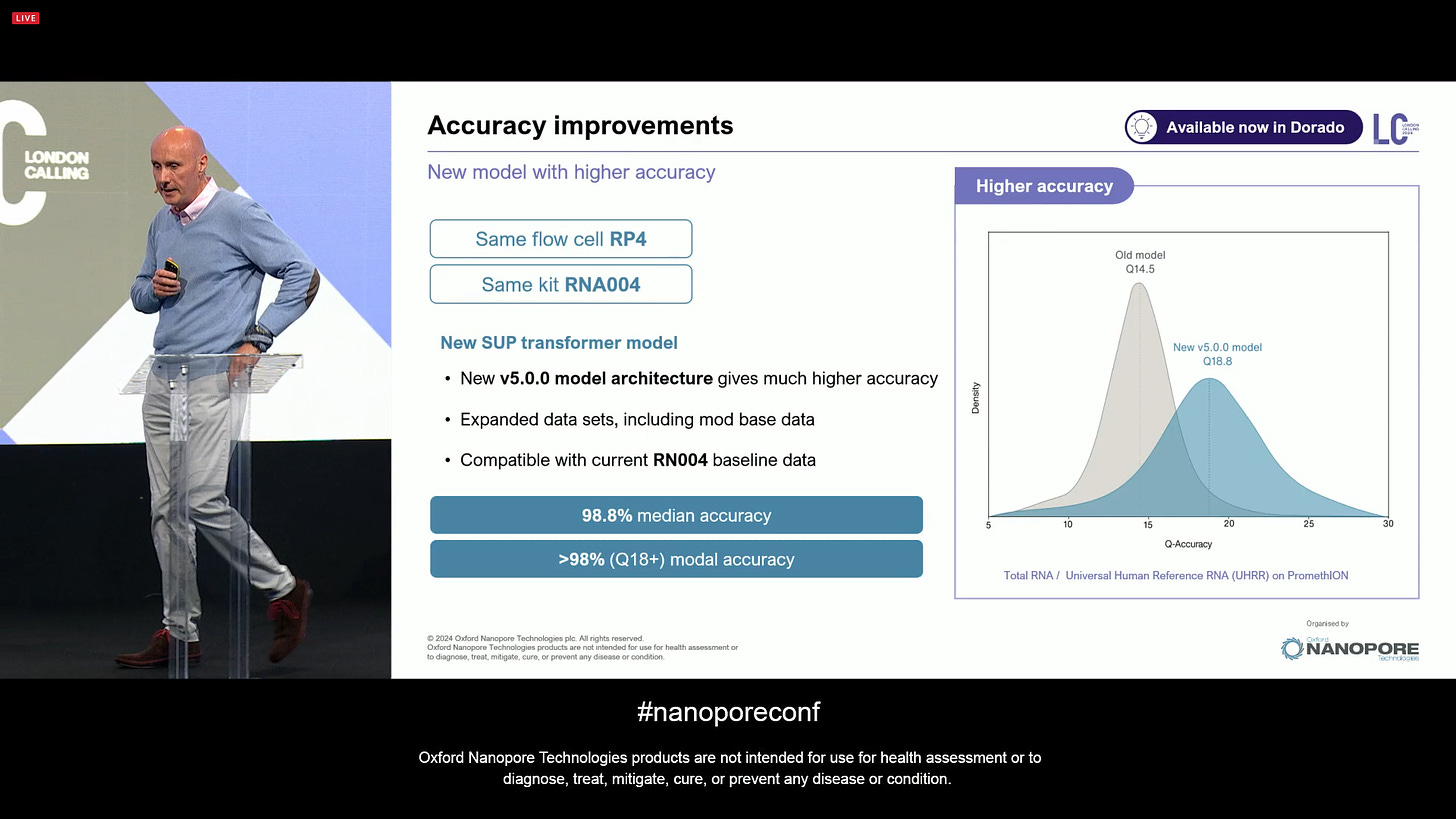
There has also been an accuracy improvement: a new v5 model architecture, including an expanded reference dataset, including mod base data, all compatible with the current RNA004 baseline data. The median accuracy is now 98.8% with Q18+ modal.
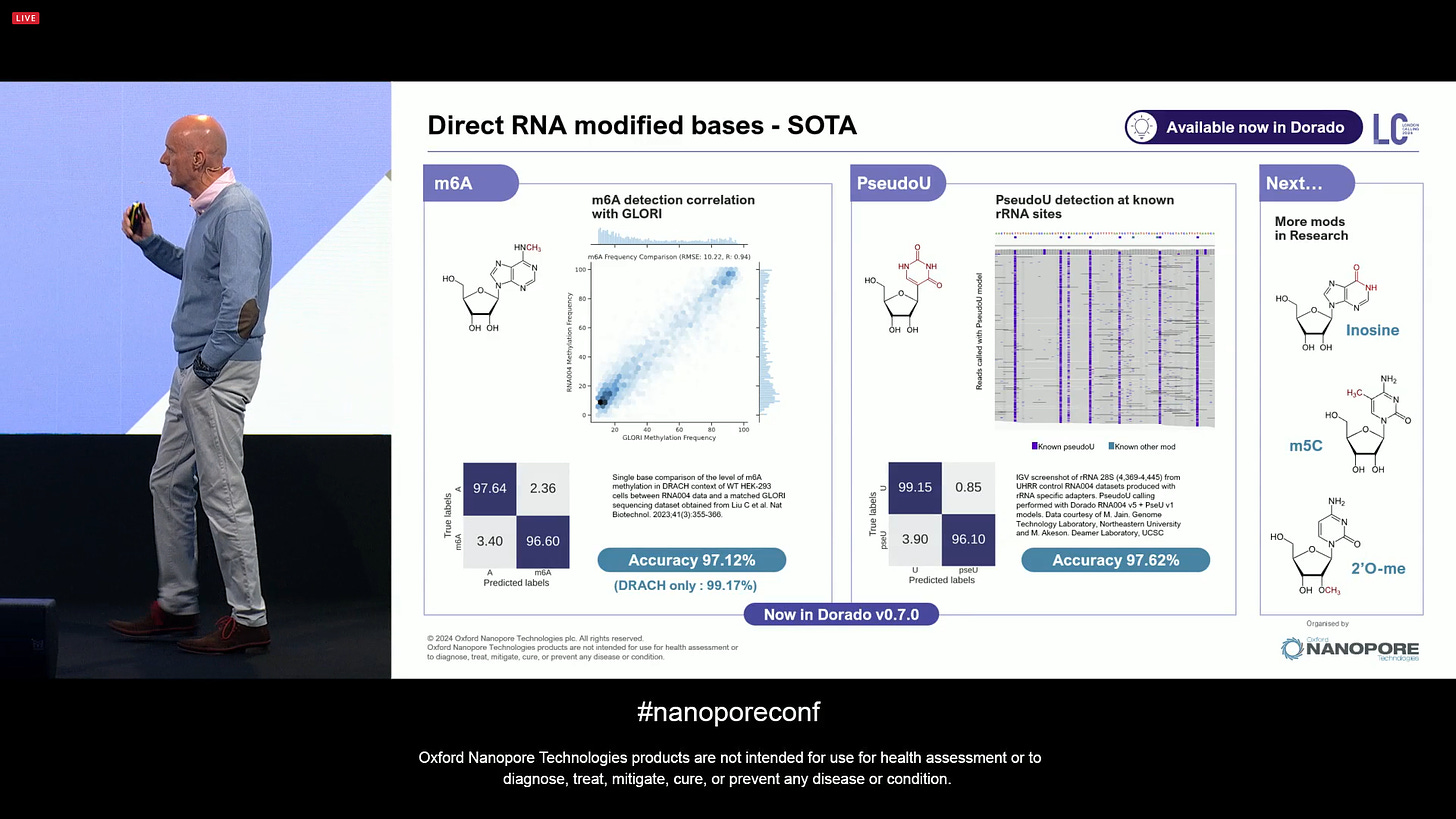
ONT can now claim SOTA on Direct RNA modified bases, all available now in Dorado v0.7. The next mods for RNA are Inosine and 2’O-me.
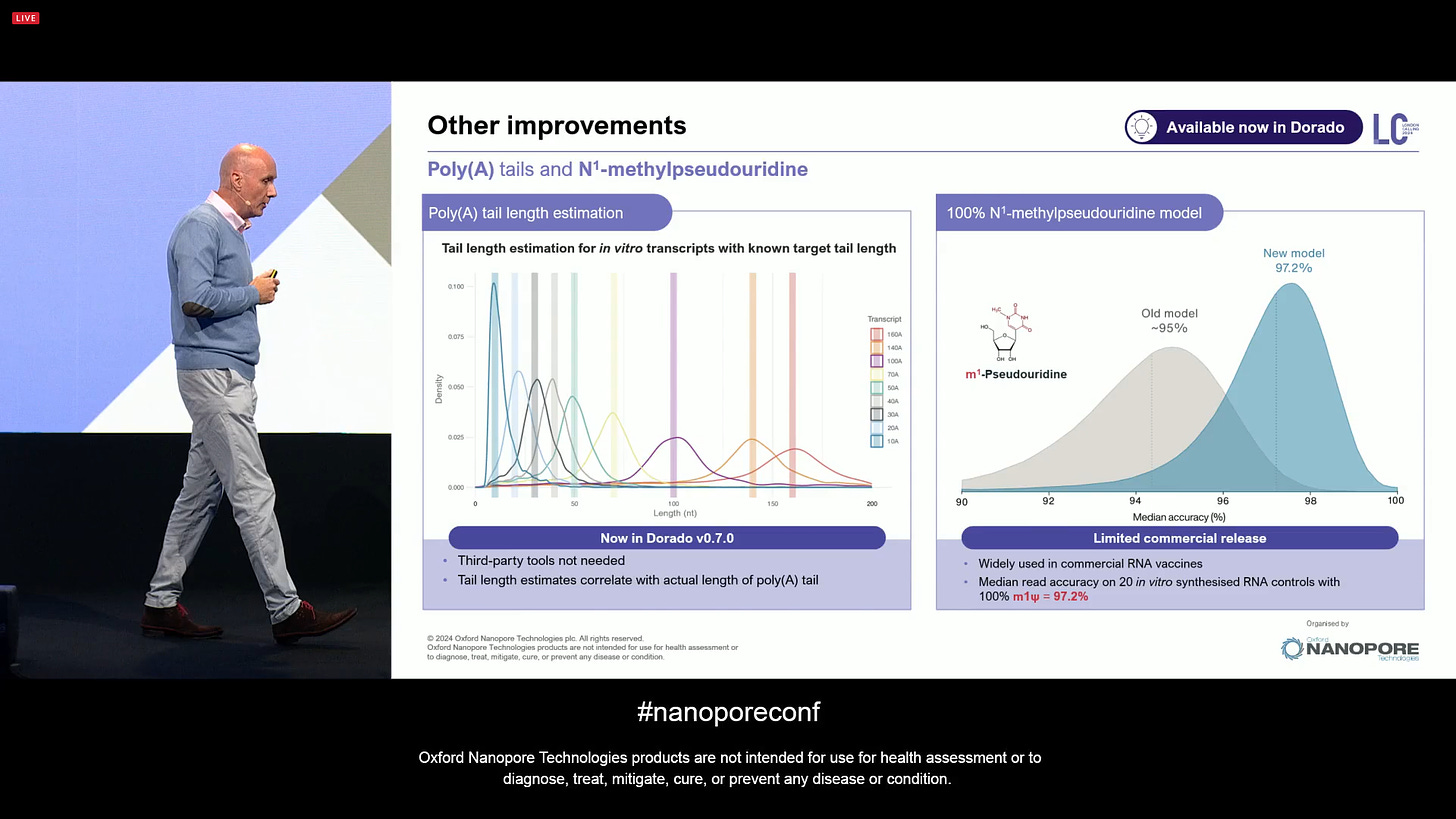
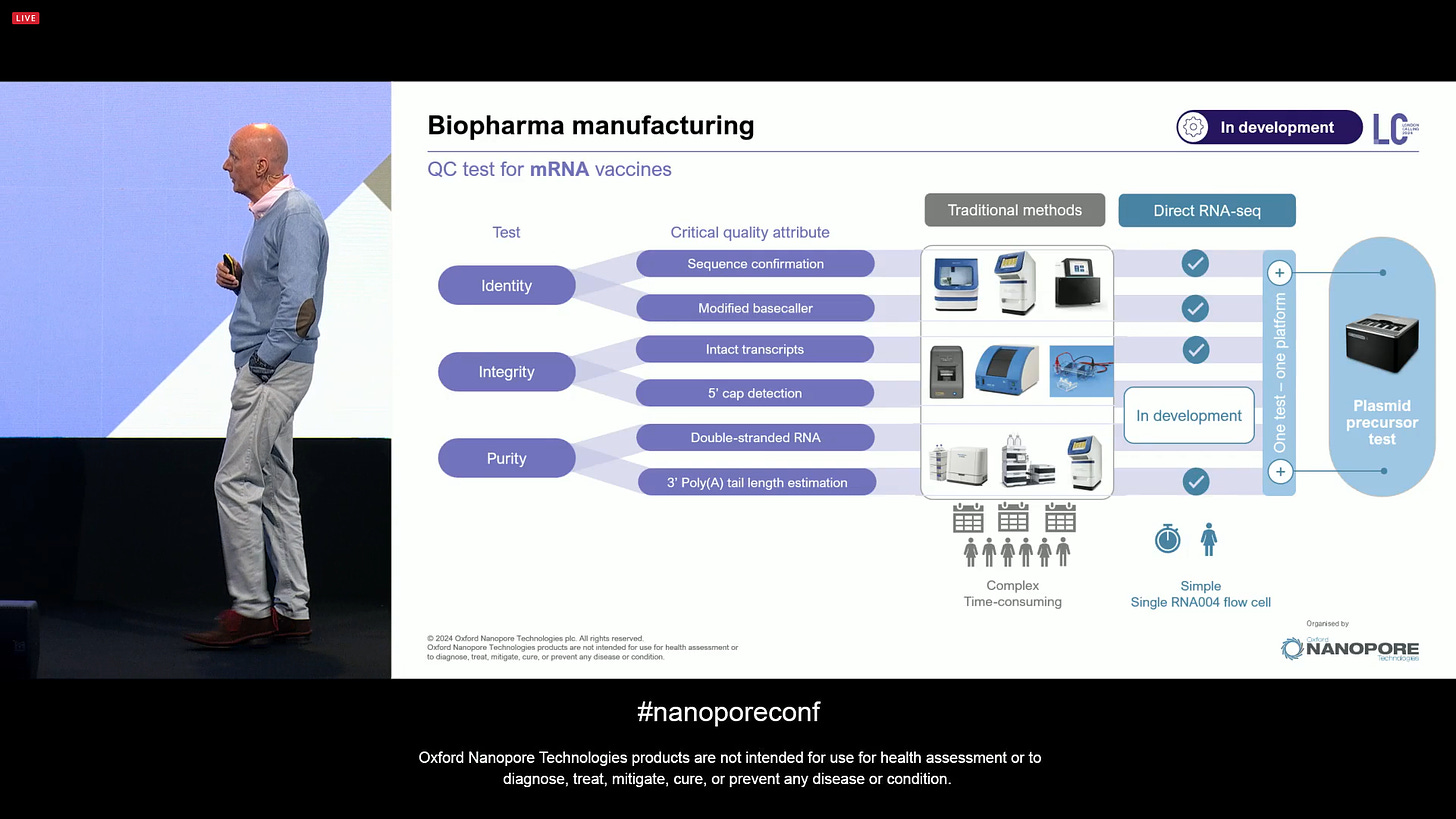
Other Direct RNA improvements include the improved estimation of poly-A lengths, and the N1-methylpseudouridine detection now at 97.2% accuracy, which is important for mRNA vaccine manufacturing and other biopharma manufacturing applications.
Indeed, the Direct RNA-seq method by ONT can now replace traditional methods that can be complex and time-consuming, and there is more in development.
Overall, Direct RNA sequencing continues to grow and mature.
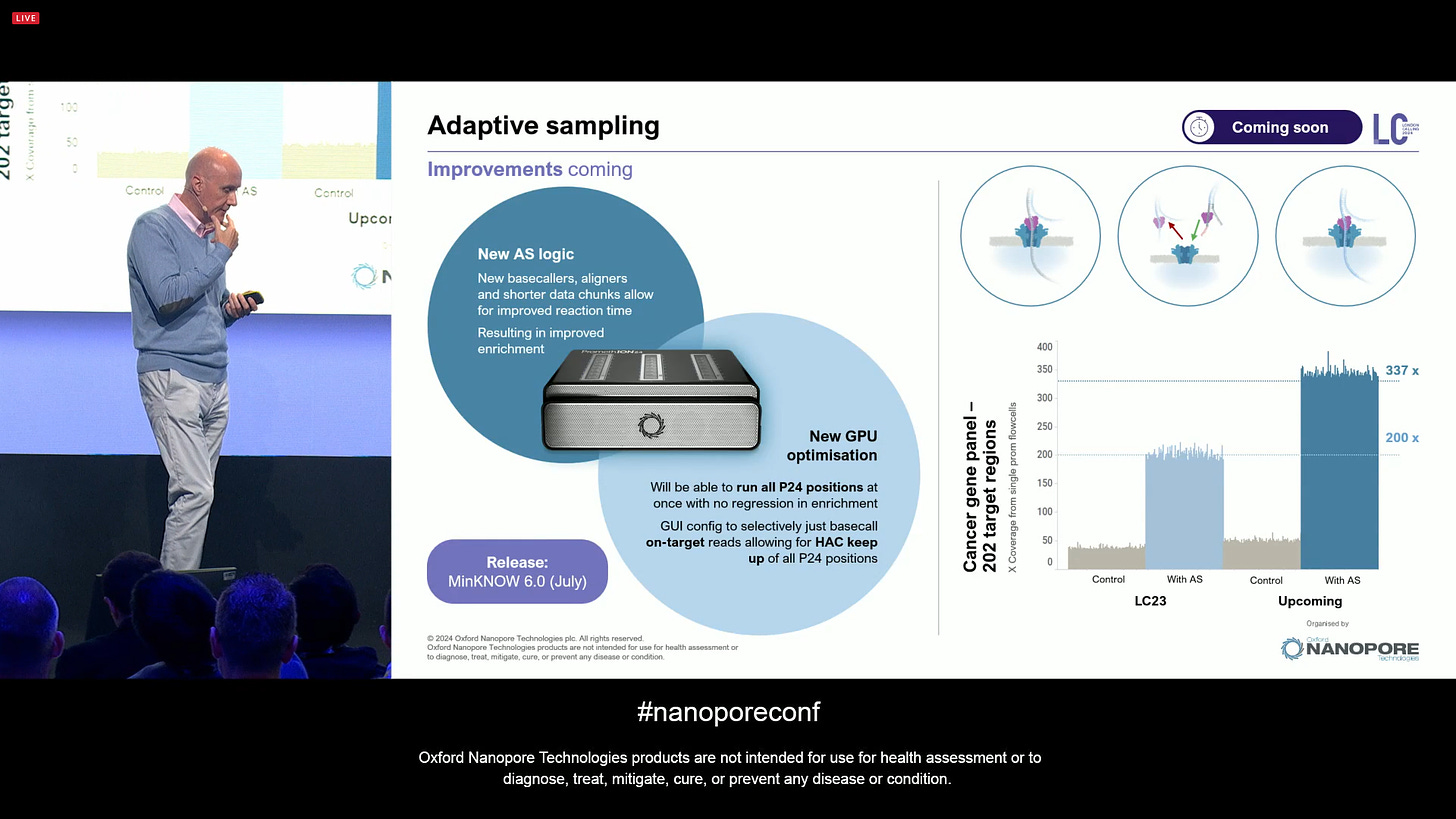
Moving on to another very special feature of ONT sequencing: adaptive sampling. There is now a new version of this feature, which will be released in MinKNOW 6.0 (July), with a new logic and new GPU optimisations, so that it can keep up with HAC basecalling of up to 24 concurrent PromethION flowcells (a full P24 run). The improvements mean that for a cancer gene panel of 202 target regions, the enriched coverage goes from 200x as it was the case in 2023 to 337x now with these updates. This also gives you “skim sequencing” of the whole genome for free, which is important to applications like CNA calling of tumour samples.
Overall, MinKNOW 6.0 is a big release, with lots of features included.
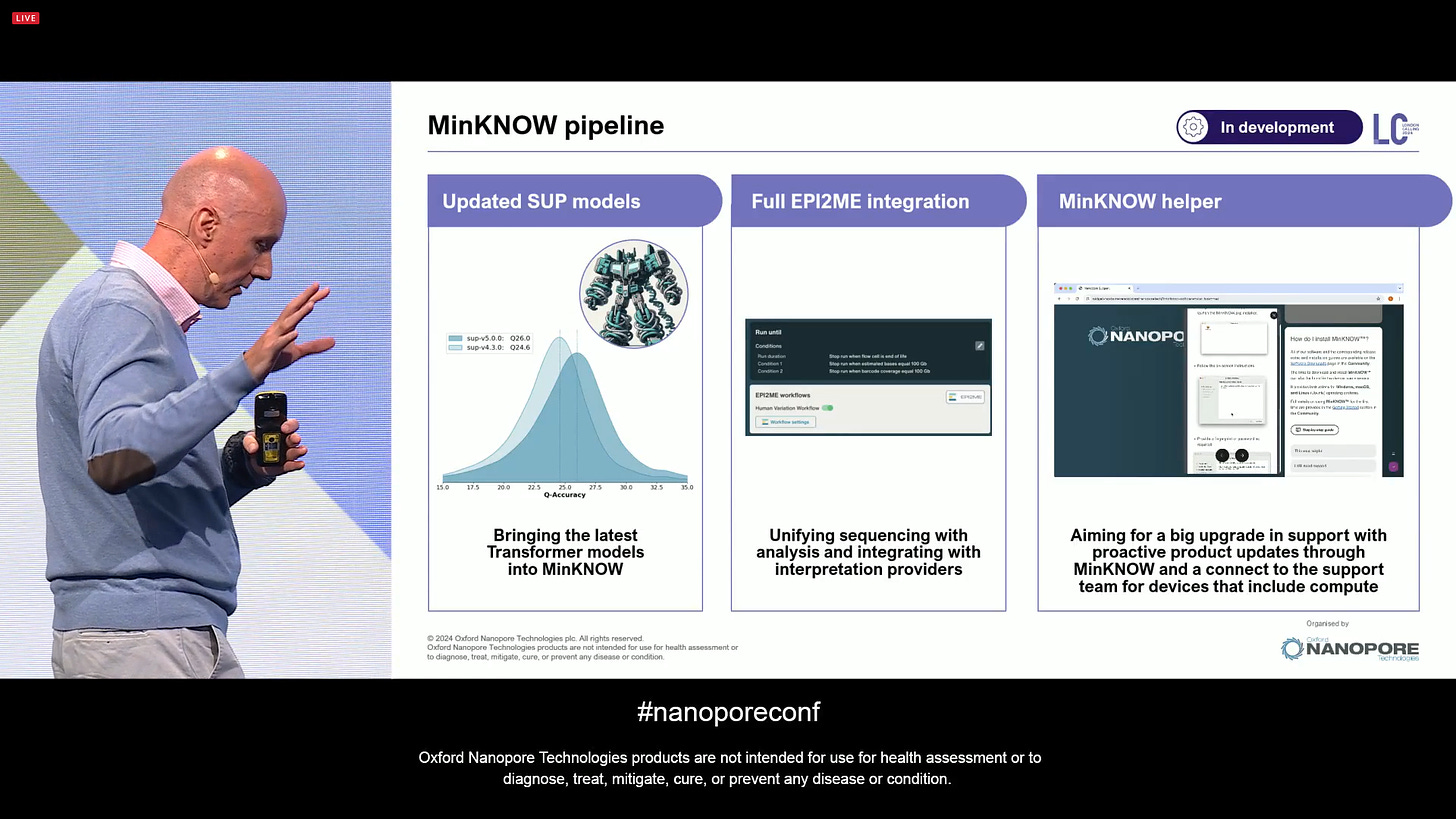
The MinKNOW software development pipeline now includes the next few pieces of work to do: updated SUP models, the Transformer-based models in dorado v0.7, a full EPI2ME integration to run locally, and direct customer support.
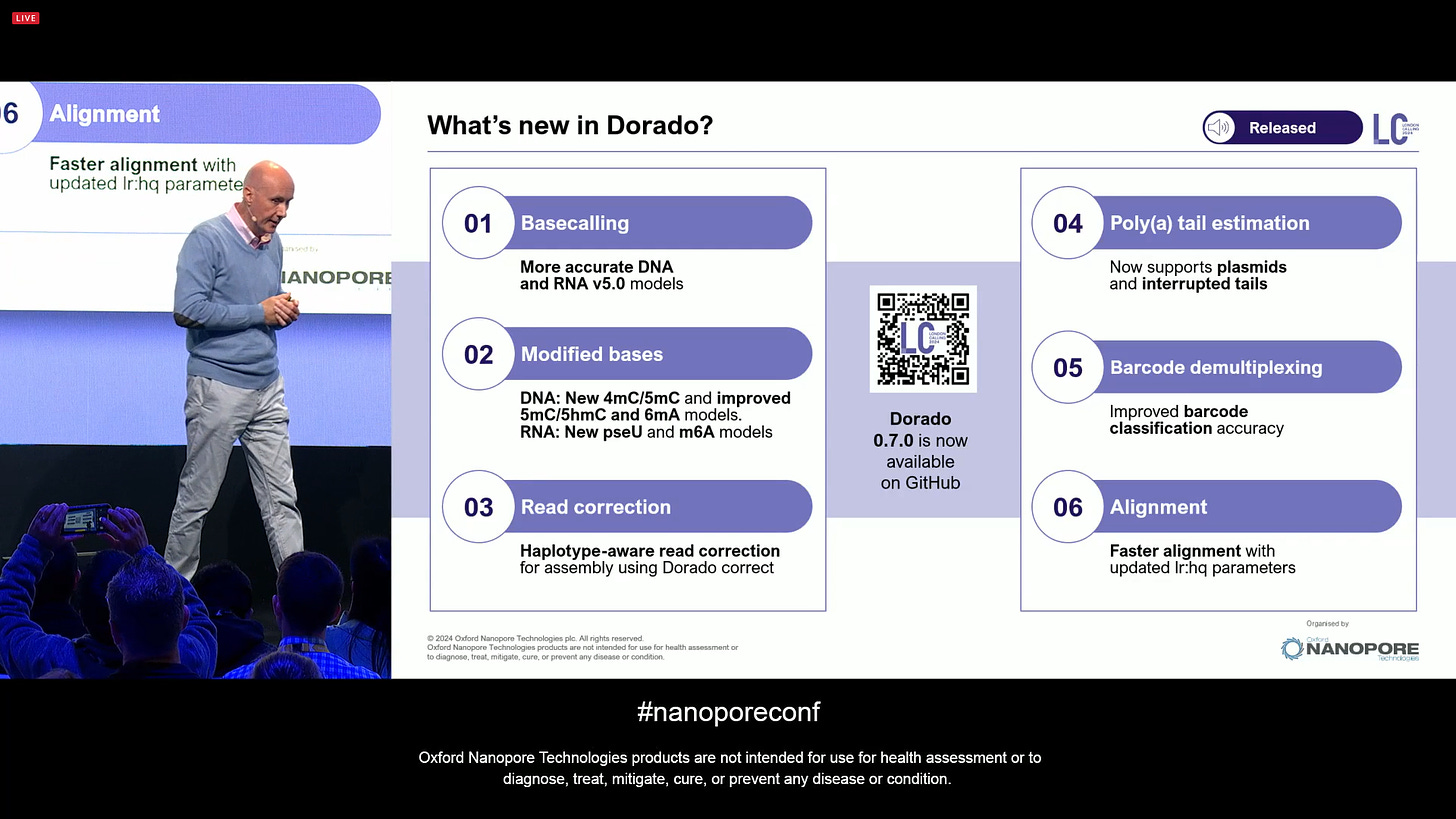
What’s new in Dorado? The basecalling is now more accurate with the v5 models, including both DNA and RNA. New DNA mods 4mC and RNA mods pseU and m6A, and improved 5mC/5hmC and 6mA models. Poly(a) estimation, barcode demultiplexing, alignment, all integrated into dorado, now available on GitHub.
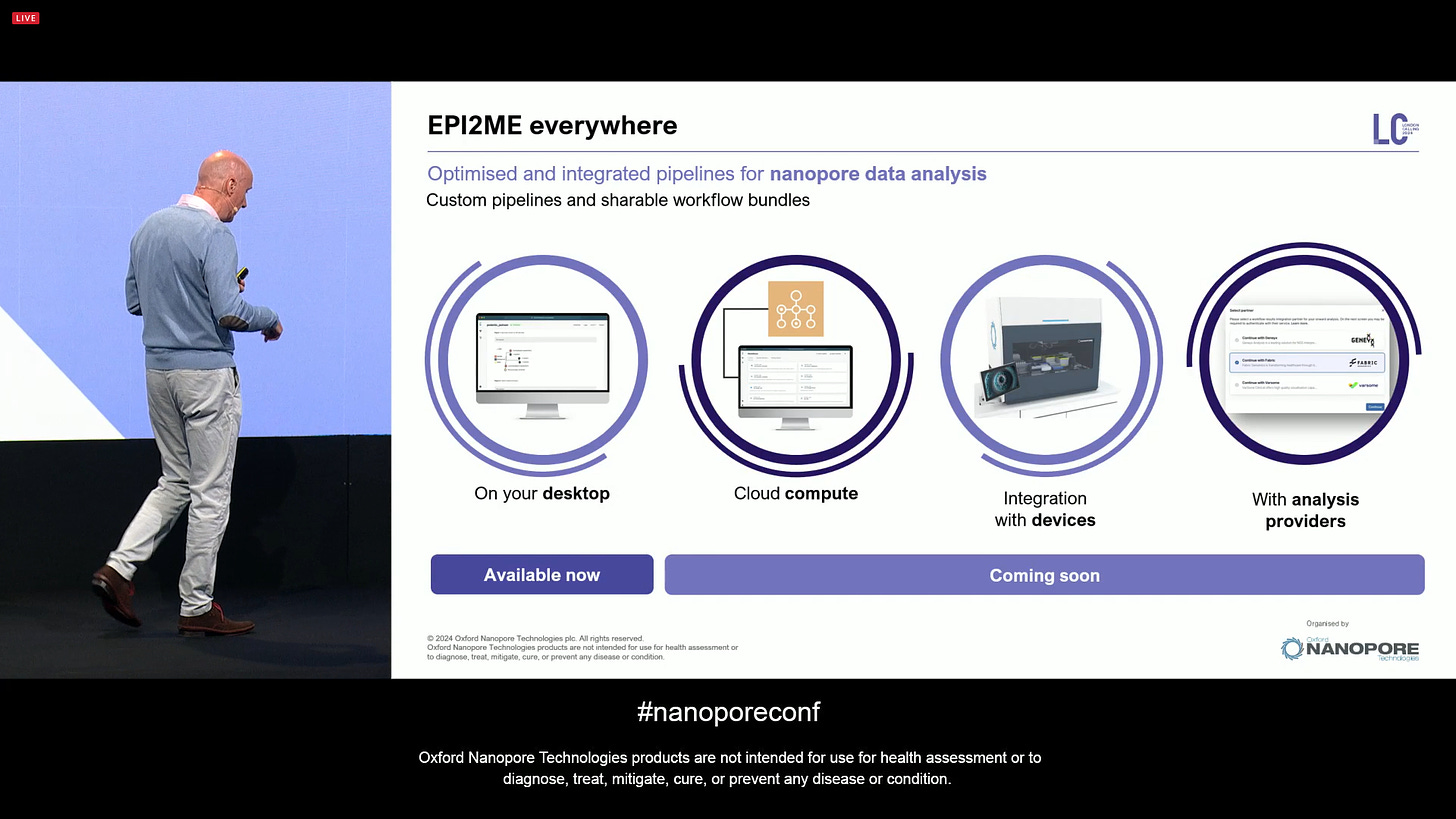
EPI2ME continues to grow and mature: you can run it on your desktop, and now working on deployment on the cloud, utilising the computational power of the instruments and incorporating more third-party providers. EPI2ME is growing nicely.
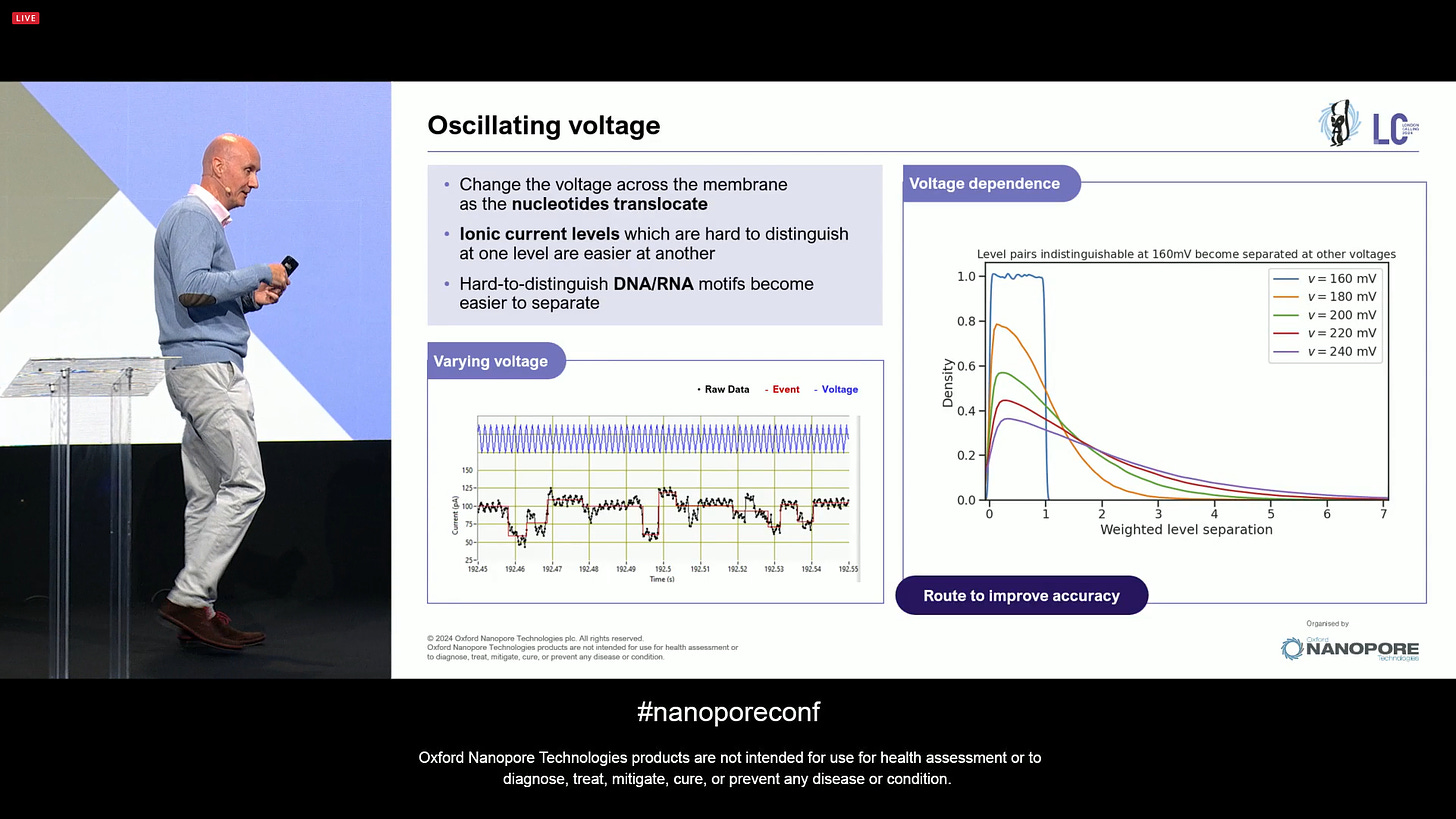
Moving towards some of the more forward-looking tech developments: one of them is the use of oscillating voltage to improve accuracy. When changing the voltage, the strands are stretched and one is able to read them more than once at different voltages, which means it can resolve hard-to-distinguish DNA and RNA motifs. There will be more news soon.
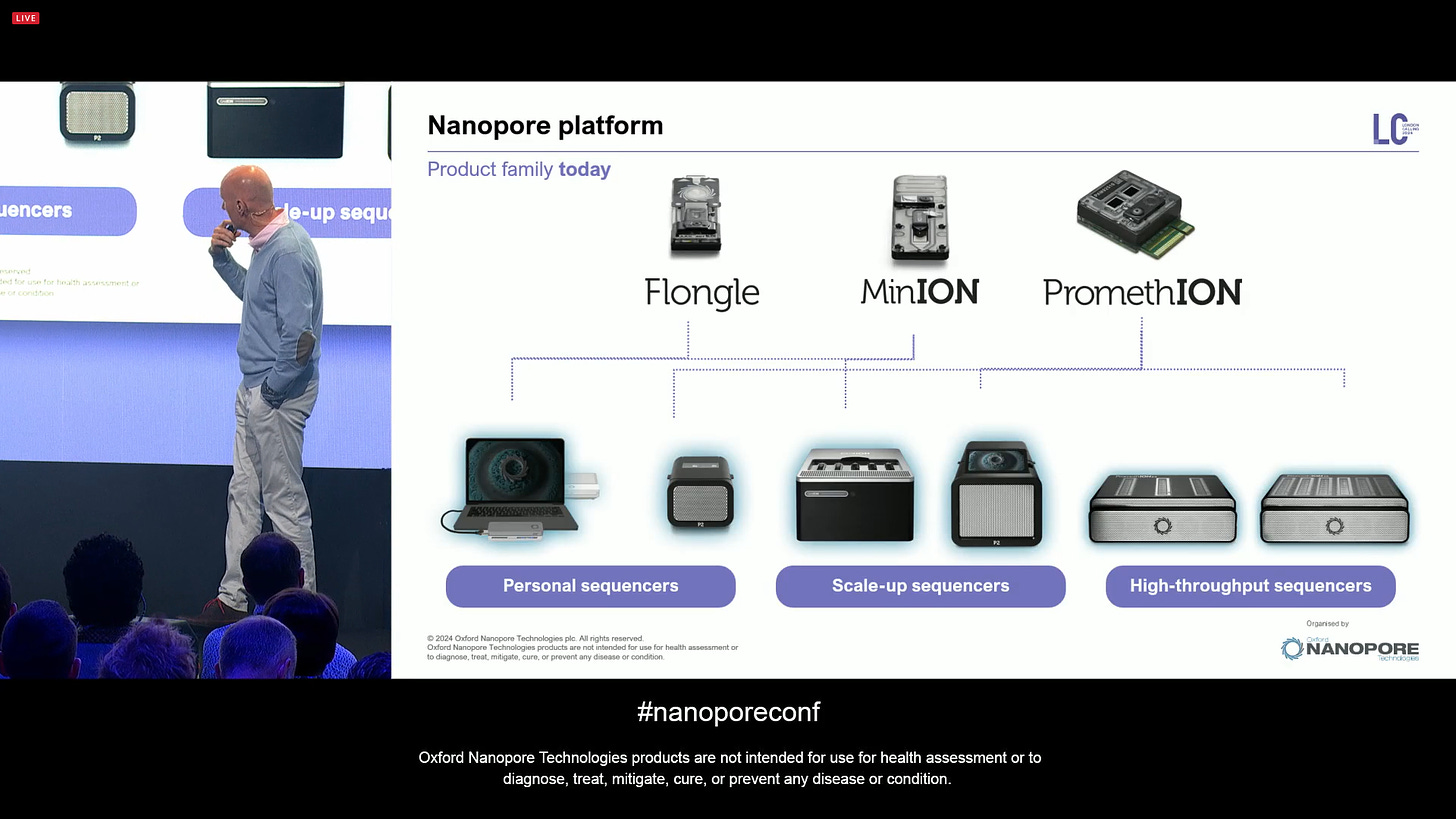
The line up of flow cells and instruments as they are. The P2 line has been very well adopted.
All devices are fully available in store, the last one is P2I. But what else is coming soon?
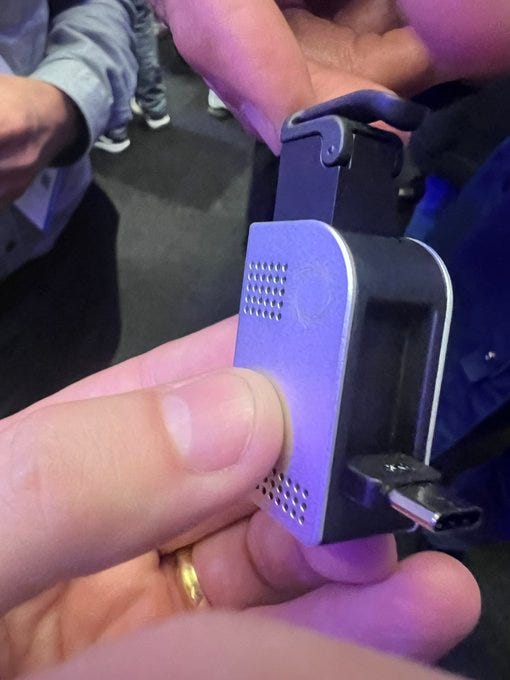
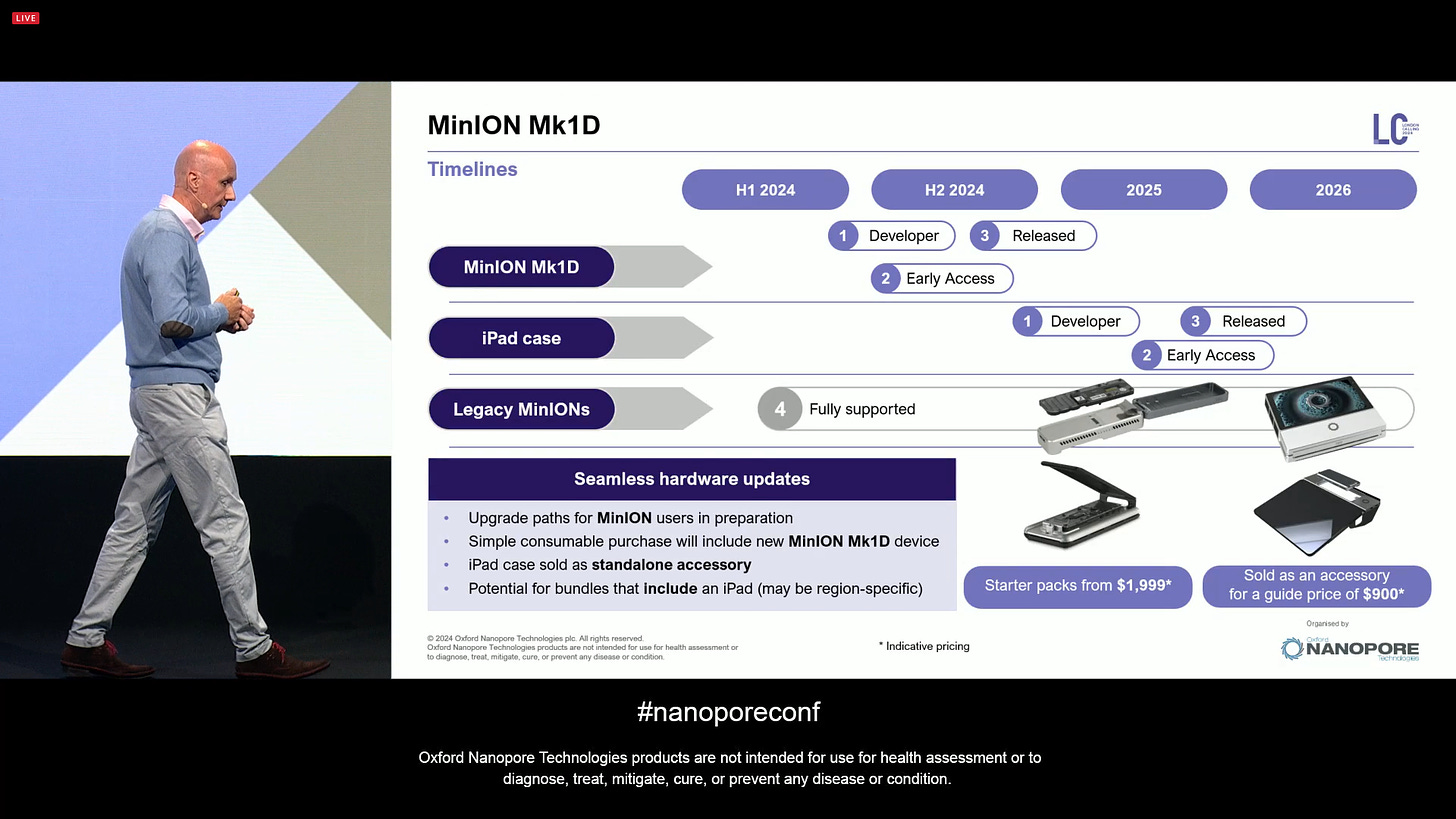
The venerable MinION will soon receive an upgrade, in the form of the Mk1D, in many ways this is similar how mobile phones get better tech inside to support more features. Here the USB-C upgrade, a better temperature control and simplified installation are some of the differences between the upcoming new MinION and the previous generation. Early Access devices will begin shipping in July and Broader access will begin in September.
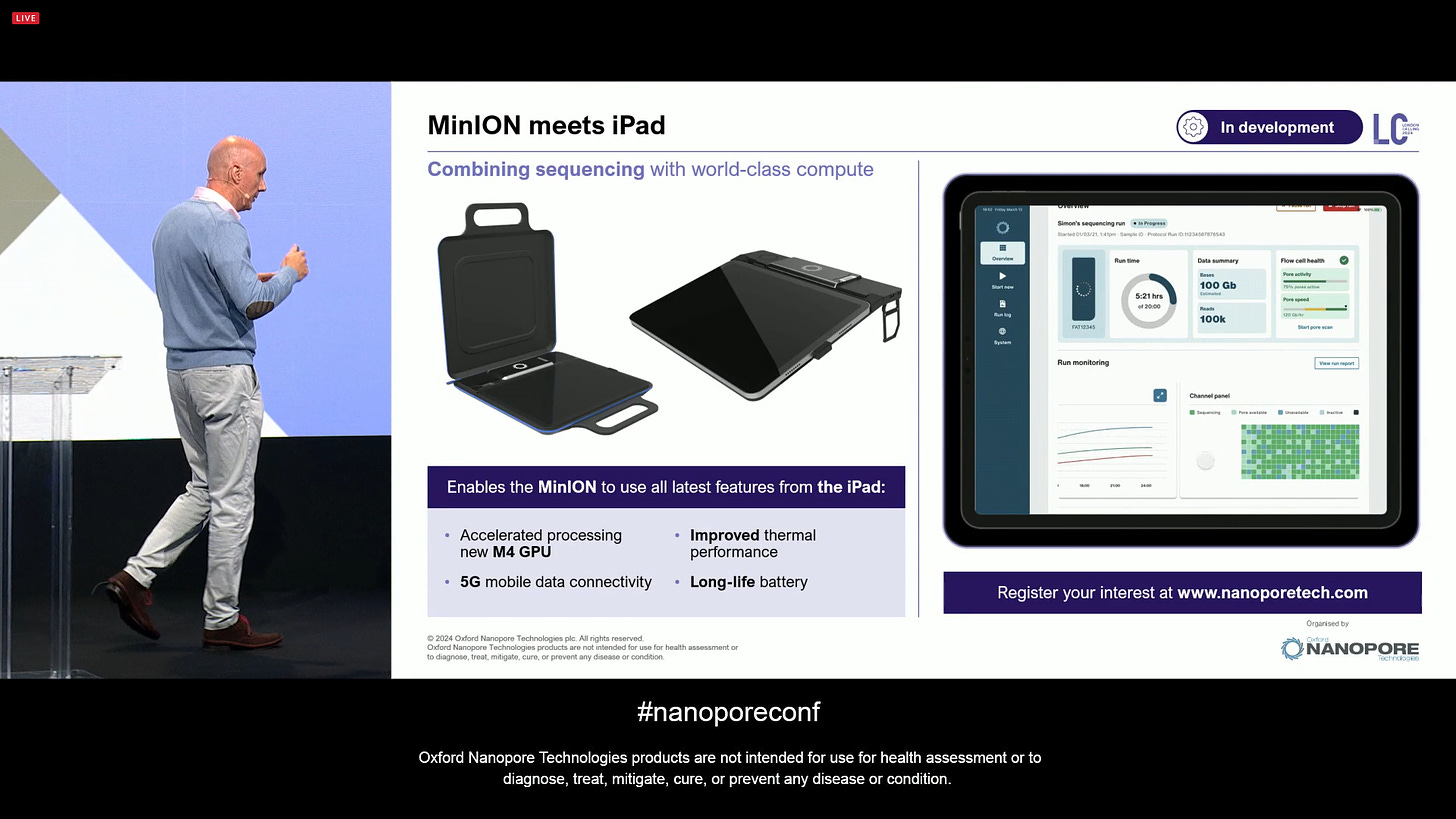
The MinION iPad is now nearing the point of release. There was a prototype on site during LC2024 for people to touch. This is predicated on the fact that the Apple Silicon chips were going to get better and better, and if ONT worked on better integrating with the devices, this could be a feature replacement of the Mk1C device, which is now showing its age.
All these will become available in the next few months, Mk1D as a $1,999 starter pack and the MinION iPad accessory for $900.
Flow cell loading is also receiving an upgrade: this will make it easier to load, a simple open and close mechanism, incorporating the light shields and good for mega long fragments, as it can now accept highly viscous samples. Prototypes during H2 2024 and release path in 2025.
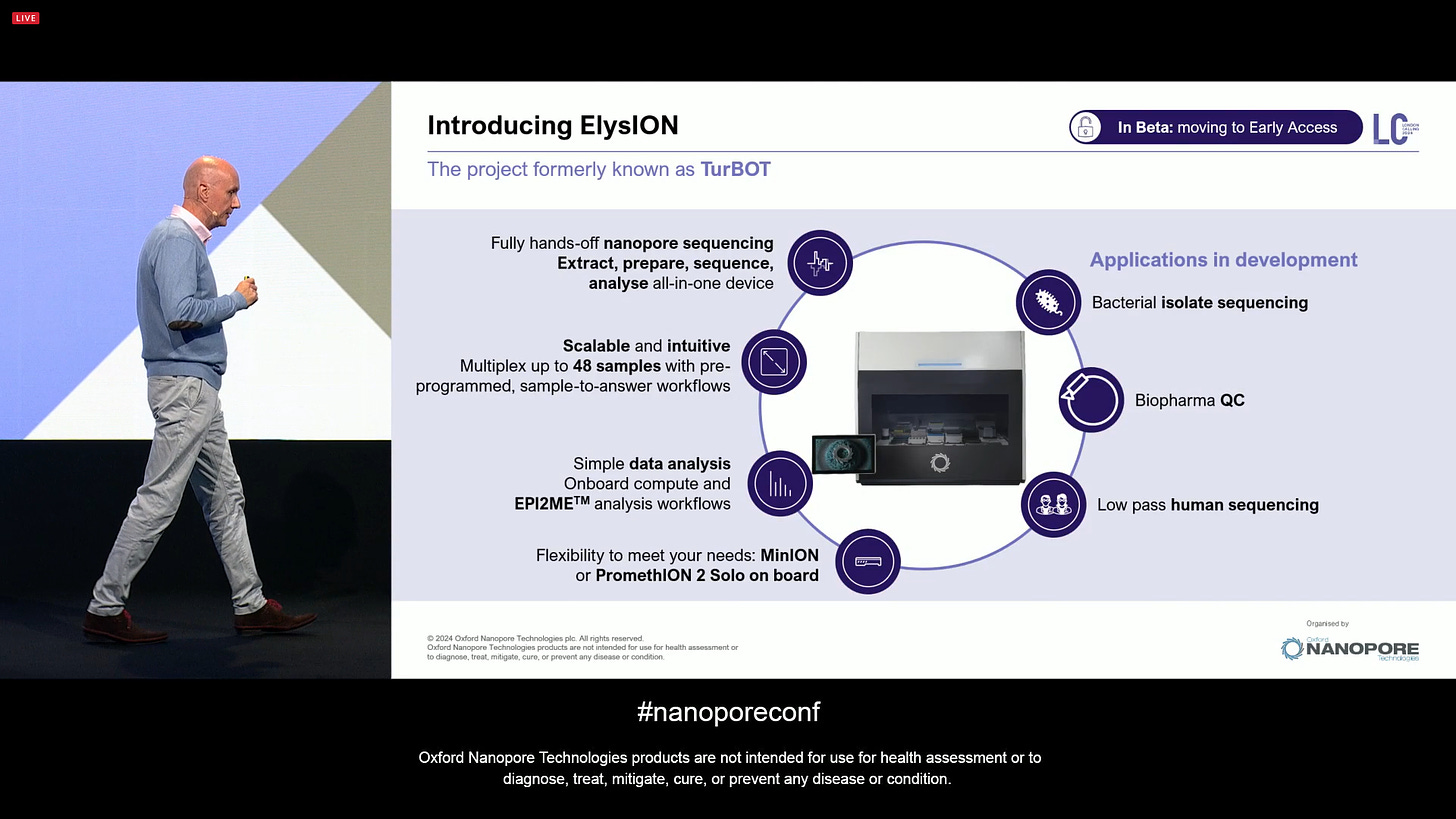
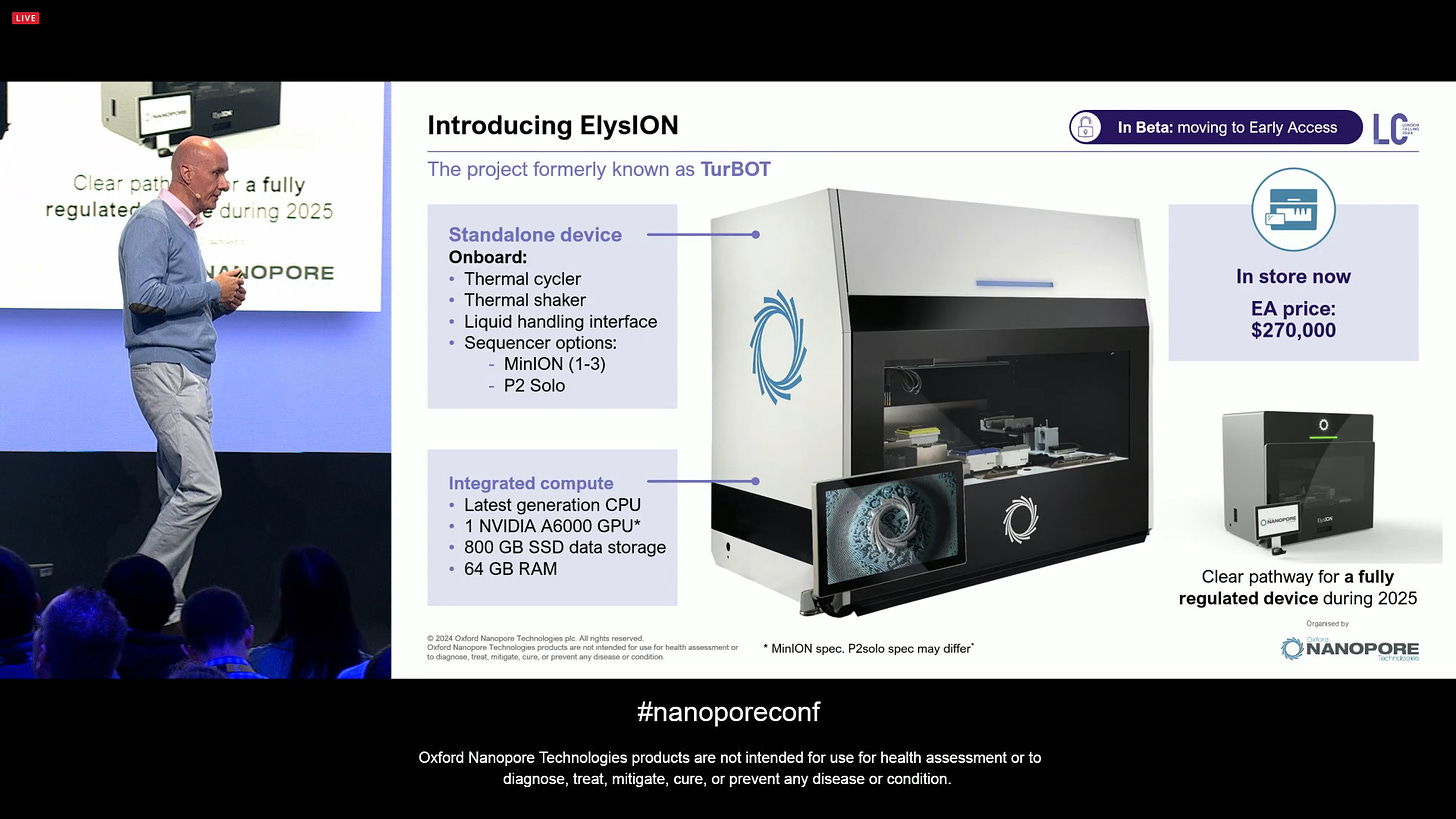
ONT introduced the ElysION (which was codenamed TurBOT until now). This is targeting a special kind of users, e.g. Biopharma QC, which want certain characteristics of the sample prep procedure that are better suited to a robotics platform.
This is now in Beta and moving to Early Access. It includes integrated compute to drive the onborad thermal cycler, shaker, liquid handling and sequencing of either MinION (1-3) and P2 Solo. It’s a bit like a Hamilton STAR but made by ONT for ONT sequencing, with an EA price of $270,000.
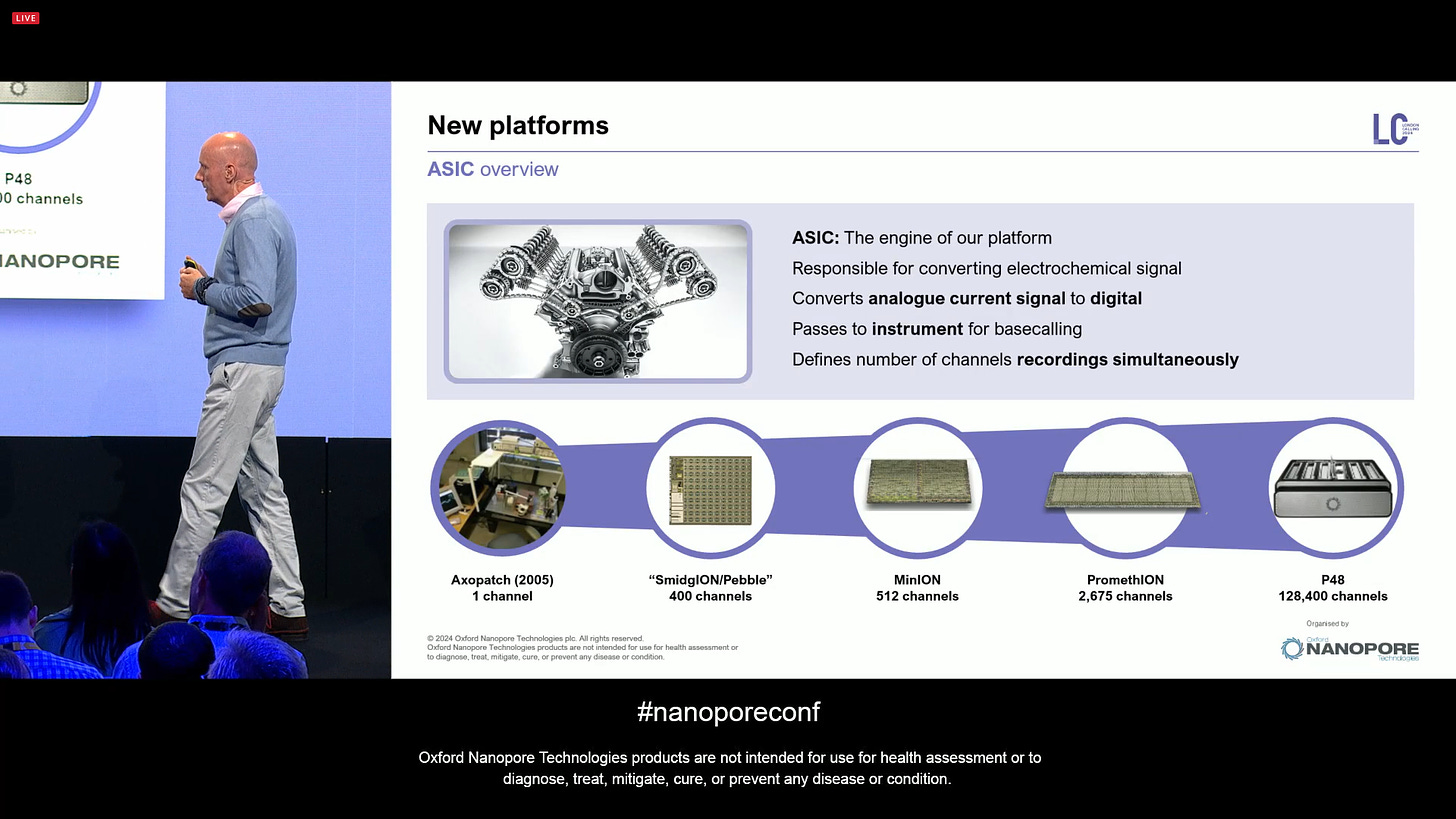
The new ASICs are going to drive new platforms for ONT. Lots of work since 2005 until now.
The SmidgION ASIC has been in development, with the following specs: 400 channels (1 mux), targeting 5-10 Gbases per flow cell, with a very small and low cost profile, and low power consumption. The flowcell chip format is similar to the MinION layout, and is now being integrated into the flow cell.
There is also a new family of instruments for the new ASIC, now under the MKII name. They will be small form factor, single flow cell instruments, with low power consumption, all from USB-C connections.
They have vertical loading, and can be “racked” in a similar fashion to the multi-lane equivalent in short-reads NGS. An 8-lane instrument would give 40-80 Gbases per run, compatible with a multichannel pipette (codename Grongle).
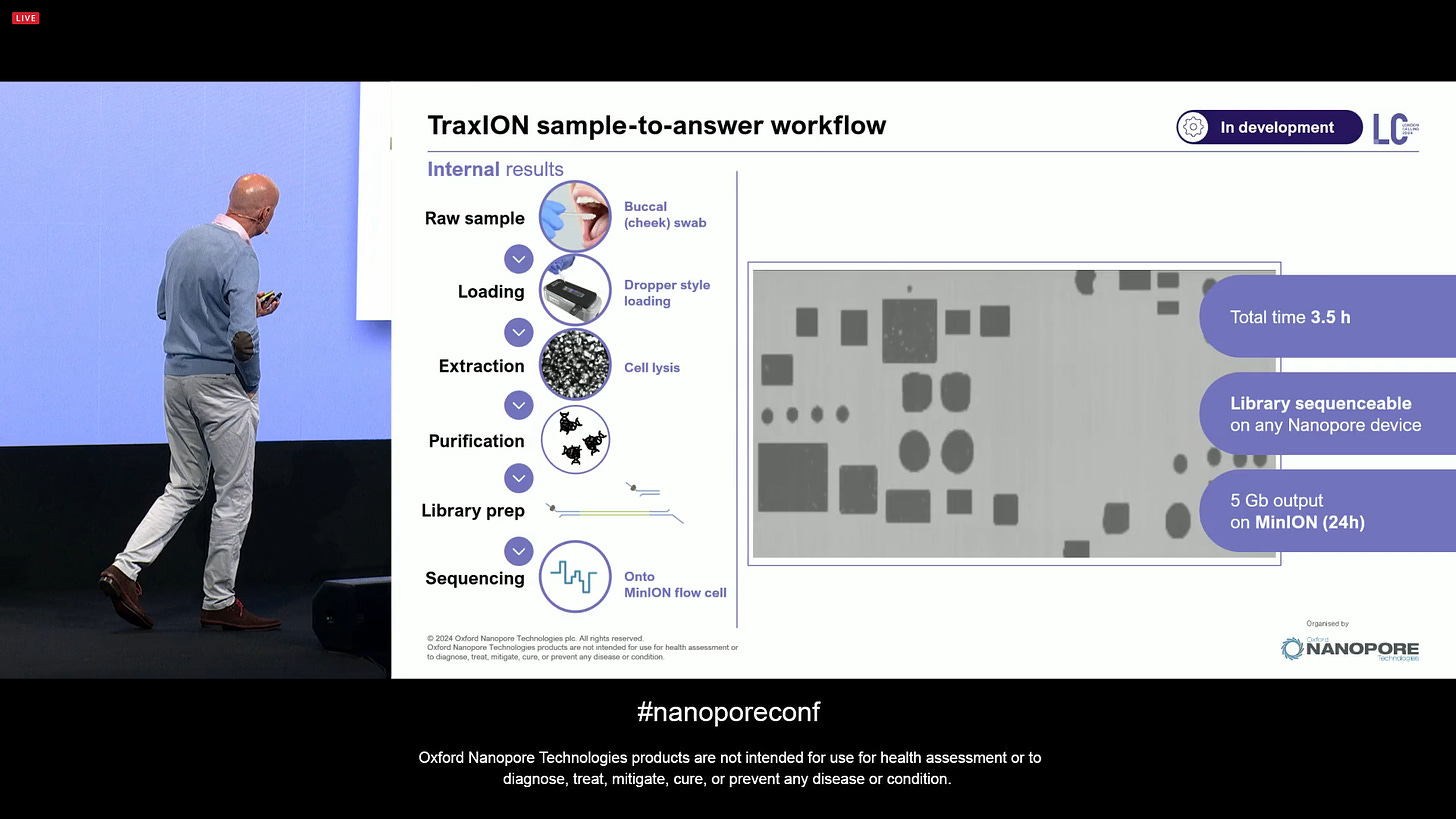
The TraxION product (previously known as a different product) is an evolution of earlier sample-to-answer device efforts. The main difference is that it now comes pre-loaded with reagents.
This means that it’s a single point of user interaction, designed so that in can have many-to-one mapping between samples and flowcells. The example here is a buccal swab, the most trial-and-tested method in the world (since COVID19). It can do cell lysis, purification, library prep, load onto the flow cell, ready to plug into the sequencer.
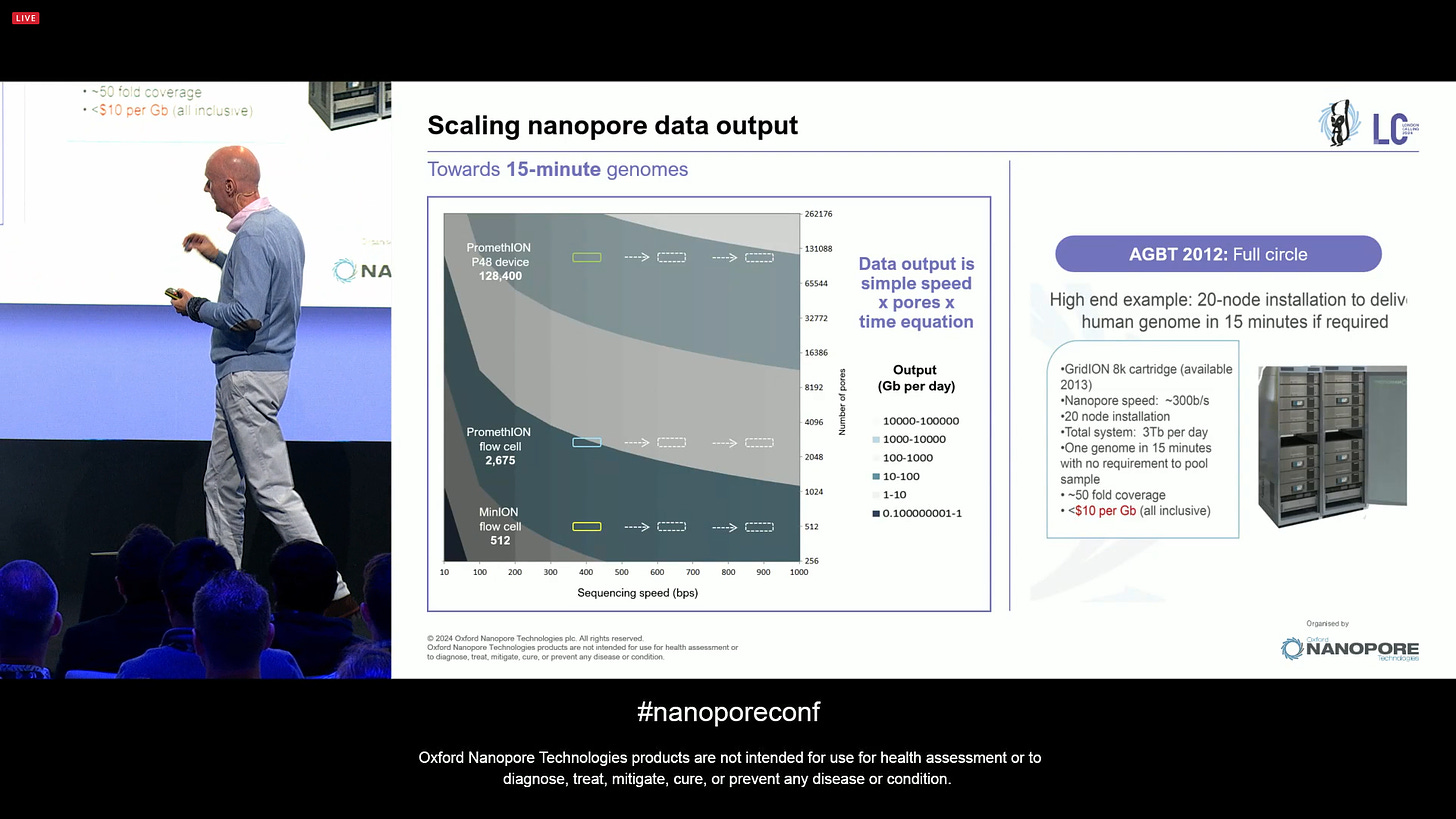
In true Steve Jobs’ style, there is “one more thing”: the new voltage sensor. Originally, this was going to be a rack of sequencers able to perform a 15 minute genome sequencing, as presented in AGBT 2012.
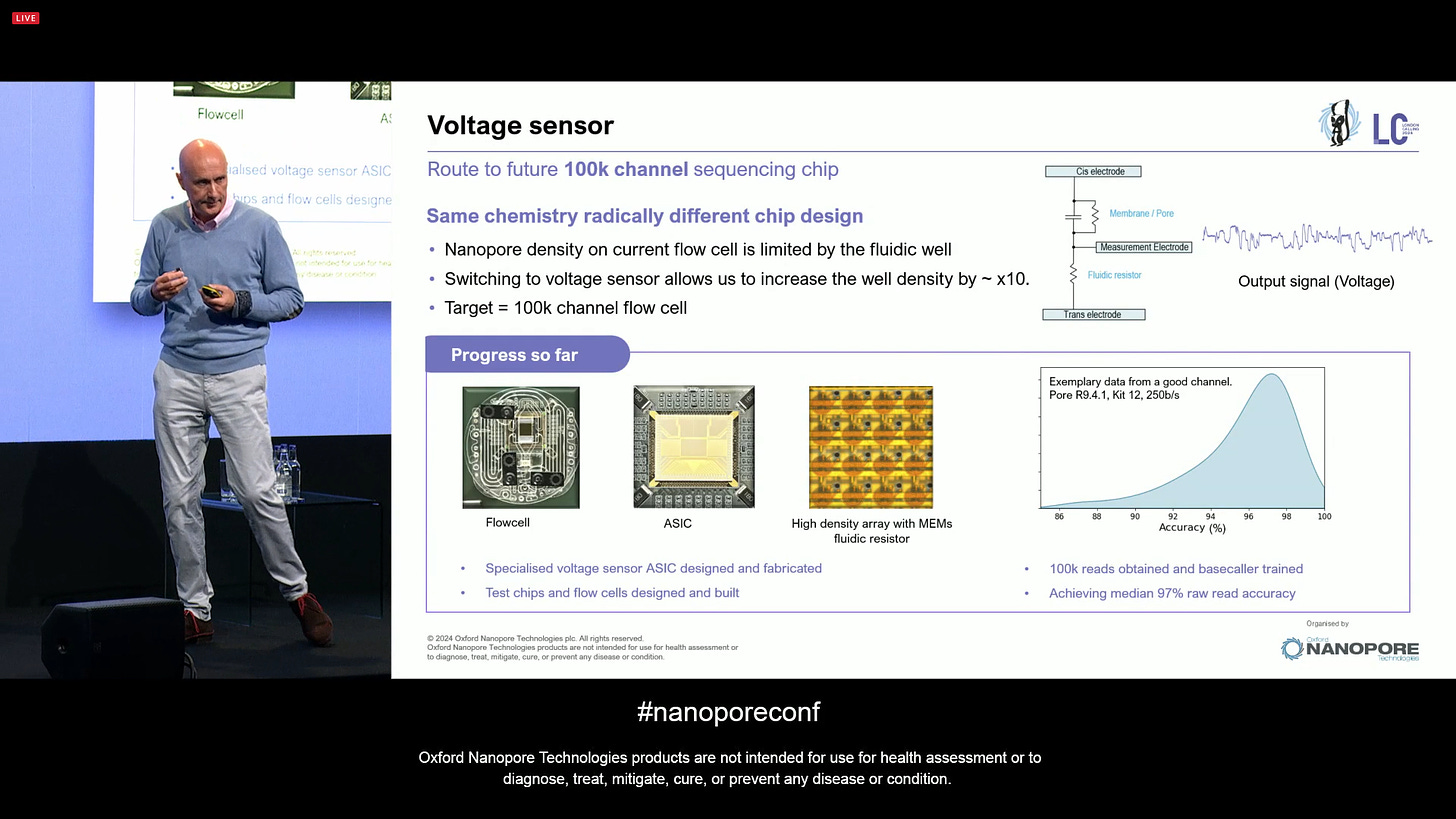
ONT is presenting the route to future 100k channel sequencing chip. This is the same chemistry but with a radically different chip design, where the density is not limited by the fluidic well but rather it utilises a voltage sensor that can increase the density by 10x. The initial target is 100k channels.
What would ONT do with such capabilities? Build a monster of a sequencer, that requires custom installation, water cooling, argon gas supply, floor reinforcement, and 24/7 onsite engineer support? This type of GargantION instrument is what is wrong with such NGS model. ONT is not like that.
Instead, this will have the shape of a shoe box, similar profile to the P2I instrument, able to use 2 flow cells of 100k channel each, delivering 10x2 Tb of data per run.
And that’s the “one more thing” that ONT is aiming at releasing soon. Clive mentioned Roche here, saying that if Roche aims for 2025+ for their Nanopore sequencer, he is confident they can release this new 100k channel sequencer sooner than Roche will do theirs.
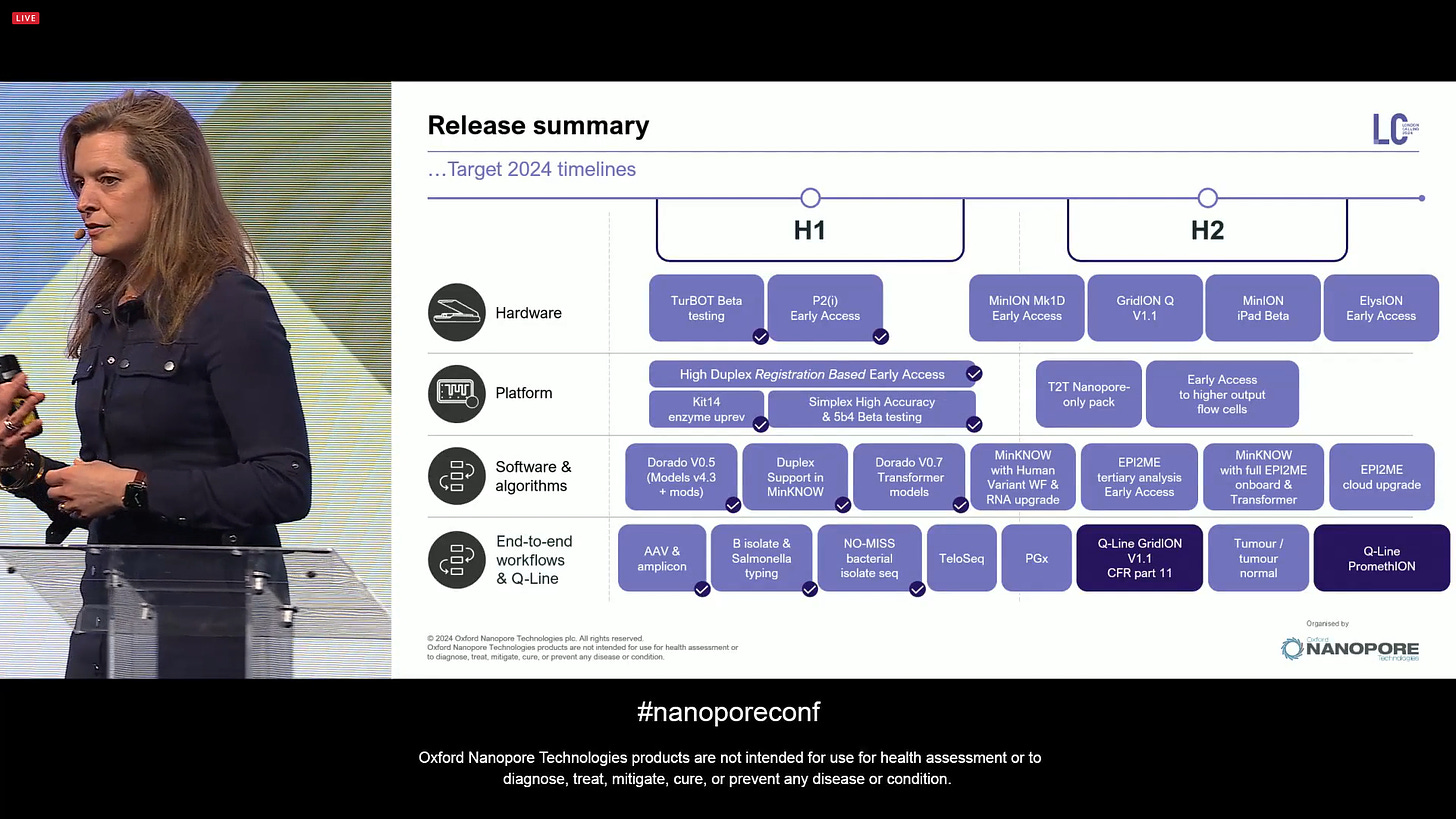
Rosemary Dokos presented the Gantt chart of releases for 2024, with a focus now on H2 both in terms of devices but also software and algorithms and associated workflows.
All in all, a jam-packed list of announcements in a session that is always exciting for people who follow Next-Generation Sequencing.